2024
Ana-Sofia Barrera

Pteropods are marine gastropods vital to pelagic ecosystems due to their importance in trophic food webs, their contribution to the ocean’s carbon cycles and fluxes, and, due to the chemistry of their shells, their role as bioindicators for ocean acidification (OA). The latter is due to how shelled pteropods build thin shells made of aragonite, a polymorph of calcium carbonate, which makes them more susceptible to dissolution in low pH conditions. Since the start of the Industrial Revolution, there’s been a massive increase in the amount of CO2 that enters our atmosphere yearly. This, in turn, has caused the pH of the ocean to drop, which has and will continue to impact the ecology of many marine species. Thus, in the last two decades, shelled pteropods and their role as bioindicators for OA have been more widely explored, given how they give insight into the impact of OA in marine ecosystems. Methods for monitoring and assessing the diversity, abundance, and community composition of pteropods in the CCE are being carried out right here at Scripps through the work of scientists like PhD candidate Anya Stajner. This work requires extensive time at the microscope and taxonomic expertise, as such many scientists are turning towards molecular biology, specifically DNA analysis, to supplement our current techniques for monitoring pteropods. Over the summer, with the guidance and mentorship of PhD candidate Anya Stajner and Professor Moira Decima, I looked into utilizing eDNA barcoding as a viable method for pteropod analysis. We aimed to create a protocol for estimating zooplankton community composition and pteropod abundance population dynamics using EtOH-preserved CalCOFI samples from the Pelagic Invertebrate Collection here at Scripps. For DNA extractions, we tested the reliability of centrifuging, and the initial results demonstrated that roughly 40% of the samples had enough DNA present for downstream applications. Likewise, we were able to develop a viable species-specific primer for L. helicina pending in-lab testing.
Ainsleigh Lloyd
 Although goose-beaked whales are common in the Southern California Bight, they can be quite elusive due to their record-breaking foraging dives. Unfortunately, goose-beaked whales are also vulnerable to noise pollution, with mass strandings and significant changes in diving behavior reported after mid-frequency active sonar (MFAS) exercises. In order to effectively conserve this species, behavioral and population density models are a necessity. This summer I worked in the Marine Bioacoustics Research Collaborative (MBARC) under the mentorship of Dr. John Hildebrand, Lauren Baggett, and Dr. Simone Baumann-Pickering. I investigated goose-beaked whale kinematics by calculating their 3D rotations using acoustic tag data. In MATLAB, I processed and annotated DTAG data to obtain raw magnetometer and accelerometer values. I then converted the magnetometer and accelerometer data into tag rotations. By calculating the rotation matrix needed to convert the tag reference frame to the whale reference frame, I was able to derive the rotations of the whale. The rotation values provide us with quantitative data on diving behavior, which can then be applied to population density estimates and pre existing models such as whale tracks.
Although goose-beaked whales are common in the Southern California Bight, they can be quite elusive due to their record-breaking foraging dives. Unfortunately, goose-beaked whales are also vulnerable to noise pollution, with mass strandings and significant changes in diving behavior reported after mid-frequency active sonar (MFAS) exercises. In order to effectively conserve this species, behavioral and population density models are a necessity. This summer I worked in the Marine Bioacoustics Research Collaborative (MBARC) under the mentorship of Dr. John Hildebrand, Lauren Baggett, and Dr. Simone Baumann-Pickering. I investigated goose-beaked whale kinematics by calculating their 3D rotations using acoustic tag data. In MATLAB, I processed and annotated DTAG data to obtain raw magnetometer and accelerometer values. I then converted the magnetometer and accelerometer data into tag rotations. By calculating the rotation matrix needed to convert the tag reference frame to the whale reference frame, I was able to derive the rotations of the whale. The rotation values provide us with quantitative data on diving behavior, which can then be applied to population density estimates and pre existing models such as whale tracks.
So far in this project, we have analyzed the diving behavior of two goose-beaked whales, using both Euler angles (pitch, roll, and azimuth) and quaternions to represent whale rotation. We have found that descent pitches are far steeper than ascent pitches. Additionally, goose-beaked whales have a varied azimuth when descending, showing a uniform or near-uniform distribution of azimuth angles, indicating that their descent may look more like a spiral than a straight diagonal path to their foraging depths. This is contrasted by ascent azimuth, which indicates a clear preference for one direction. However, since this project is still in the early stages, more analysis and a greater sample size are needed.
I’d like to thank Dr. Hildebrand, Lauren Baggett, and Dr. Baumann-Pickering and for providing me the opportunity to return to MBARC for my second summer, and for their invaluable mentorship. I am truly grateful for them and for the rest of the MBARC lab members who have made this an incredible experience.
Lana Lubecke

Soleil Michaud
 Small crustacean plankton, such as copepods, provide food, refuge, and transportation to hitchhiking bacteria. The nutrient-rich exoskeletons of these organisms create an ideal environment for the bacteria to thrive, and warming oceans may intensify these dynamics, threatening public health and food security. As the global sea surface temperatures continue to rise, evidence has shown that Vibrio related infections have increased in humans. Marine heat waves have also been associated with Vibrio-induced mass mortality of aquaculture species. As sea surface temperature continues to increase with global warming, Vibrio infections pose a significant and growing threat to global health. Under the guidance of Dr. Julie Dinasquet, we have worked to develop a protocol for extracting DNA from formalin-preserved samples with the aim to investigate the long-term population dynamics of Vibrio spp. for the first time. Using CalCOFI’s unique and expansive 75-year archive of formalin-preserved zooplankton from Southern California. We plan to build a time series of the population of the opportunistic marine pathogen Vibrio spp. dynamics in Southern California.
Small crustacean plankton, such as copepods, provide food, refuge, and transportation to hitchhiking bacteria. The nutrient-rich exoskeletons of these organisms create an ideal environment for the bacteria to thrive, and warming oceans may intensify these dynamics, threatening public health and food security. As the global sea surface temperatures continue to rise, evidence has shown that Vibrio related infections have increased in humans. Marine heat waves have also been associated with Vibrio-induced mass mortality of aquaculture species. As sea surface temperature continues to increase with global warming, Vibrio infections pose a significant and growing threat to global health. Under the guidance of Dr. Julie Dinasquet, we have worked to develop a protocol for extracting DNA from formalin-preserved samples with the aim to investigate the long-term population dynamics of Vibrio spp. for the first time. Using CalCOFI’s unique and expansive 75-year archive of formalin-preserved zooplankton from Southern California. We plan to build a time series of the population of the opportunistic marine pathogen Vibrio spp. dynamics in Southern California.
Formalin-preserved samples have been historically considered inaccessible due to the impact of formalin on DNA degradation, protein cross linking and PCR inhibition. Despite these challenges, several recent studies have successfully recovered DNA from formalin-preserved samples. The first part of this REU project was to assess if DNA can be successfully extracted from formalin-preserved Calanus pacificus samples, before investigating the bacterial population associated with the copepod.
Samples were obtained from the Pelagic Invertebrate Collection from 1984-2024. All of these samples were collected on the quarterly California Cooperative Oceanic Fisheries Investigation (CalCOFI) cruises.
The project focuses on Calanus pacificus, a key copepod in the California Ecosystem.C. pacificus, is the dominant copepod species in coastal Southern California. It is an important food source for fishery species and thus a likely vector of pathogens. I was trained to sort and collect several single C. pacificus specimens from each zooplankton sample.
My work so far has successfully demonstrated that DNA from both C. pacificus and Vibrio can be extracted from these preserved samples dating back to 1984. For more recent samples DNA was recovered from single copepod organisms, and showed the presence of about 3000 Vibrio/ copepods in 2018.
Moving forward, my research will focus on optimizing the DNA extraction protocol to work on older samples, ideally beyond the current 40-year limit. I will also sequence the microbiome and Vibrio populations from these historical samples to create a comprehensive genetic database, providing a baseline for understanding the long-term ecology of copepods and their pathogens. Additionally, I hope to expand the study to include other preserved organisms, potentially building a genetic time series to track ecological changes across various marine species over time.
Abigail Severance

Over this summer, I had the privilege of working in Dr. Barbeau’s lab and researching iron forms and availability in the ocean, allowing for a greater understanding of marine photosynthesis, nutrient availability, and overall ocean environments.
Dissolvable iron in the ocean is necessary for marine photosynthesis, thereby relating iron levels to carbon uptake and overall global climate. This is most clearly illustrated in high nutrient, low chlorophyll regions, in which macronutrients are plentiful but there’s little evidence of primary productivity; here, the addition of iron stimulates the usage of the nutrients and leads to a photosynthetic bloom. Critically, the form in which this iron is present – organic, inorganic, bound, free, etc. – determines its bioavailability for photosynthesis. 99% of the dissolved iron in the ocean is bound by organic ligands of different strengths, so one of our best measures of bioavailability is measuring the relative strength of these ligands. We did this with a process known as CLE-AdCSV and surface level seawater samples from a summer 2023 cruise taken off the coast of Cape Blanco in Oregon.
CLE-AdCSV refers to a competitive ligand exchange followed by adsorptive cathodic stripping voltammetry. Additions of salicylaldoxime, a known, electroactive ligand, are allowed to compete with natural seawater ligands for Fe complexation. Electrochemical measurements of the Fe-SA complex allow us to determine the strength and concentration of the natural ligands. After preparing aliquots of the samples – including additions of boric acid to buffer the samples to seawater’s natural pH and iron additions of varying concentrations to bind to all ligands within the sample – an electroactive complex is formed which adsorbs to the mercury drop cathode used. A titration curve is produced from running a voltage sweep over this sample, and from this curve, further processing allows us to determine ligand concentrations, strengths, and free iron levels in various areas.
I’d like to thank Dr. Barbeau and Max Fenton, as well as all the members of the Barbeau Lab, for all their guidance and project help this summer, it’s been an absolute pleasure to learn from everyone there!
Tongfei Yang
 Over the summer, I worked at the Semmens Lab under the guidance of graduate student Max Titcomb and Dr. Brice Semmens on the Grouper Moon project. My focus was on training AI models to create a curated dataset for face identification and pattern recognition of Nassau groupers.
Over the summer, I worked at the Semmens Lab under the guidance of graduate student Max Titcomb and Dr. Brice Semmens on the Grouper Moon project. My focus was on training AI models to create a curated dataset for face identification and pattern recognition of Nassau groupers.
A key aspect of my work involved automating fish detection in underwater videos captured by GoPro cameras. The goal was to train a model capable of placing bounding boxes around the fish in these videos to facilitate further analysis. I utilized the YOLOv9 (You Only Look Once) model, a real-time object detection algorithm. I imported the model architecture and trained it on our custom fish dataset. The first step was to label the dataset by manually placing bounding boxes on fish in video frames. After preprocessing the labeled dataset and addressing various issues, we trained a model that gave us a solid baseline. However, the model's performance declined when the number of fish in an image increased. To address this, the next step is to enhance the model's performance by integrating more training data, particularly those with multiple fish instances.
Another aspect of my work involved fish segmentation using Fishial. By feeding individual fish images from the detection model into Fishial, we obtained masks that outlined each fish. This provided us with the fish's morphology, helping with frame selection for pattern recognition.
I'm grateful for the opportunity to participate in the CCE-LTER program. The connections and experience I gained during my summer as an REU have been invaluable as I continue my academic journey and pursue future careers. I would like to thank Dr. Brice Semmens for bringing me into this incredible project, as well as the graduate students who supported me along the way: Max Titcomb, Christopher Crutchfield, and Sean Perry.
Hannah Yek
 This summer under the guidance of the Ohman Lab, including Visiting Scholar Dr. Jeff Ellen, I investigated the presence of Copepod Thin Layers in the Northwestern Mediterranean Sea. Copepod Thin Layers are difficult to measure with conventional sampling technology, but are thought to be important 'hot spots' of enhanced grazing activity on phytoplankton and elevated predation rates by copepod predators. My research focused on analyzing data from a Zooglider mission conducted in three distinct regions off the eastern coast of Spain. During the 39 day mission, the glider executed 352 dives during which 395k images were collected. After the cruise, this data was processed into 1.39M labels of individual plankters labeled by a machine learning algorithm with concurrent CTD measurements. Access to this metadata was provided via a PostgreSQL database. To facilitate this analysis, I utilized pgAdmin for preliminary queries and managing the extensive dataset generated by the Zooglider. For backend processing, I employed Python and SQLAlchemy, while Plotly and Dash were used to create interactive visualizations.
This summer under the guidance of the Ohman Lab, including Visiting Scholar Dr. Jeff Ellen, I investigated the presence of Copepod Thin Layers in the Northwestern Mediterranean Sea. Copepod Thin Layers are difficult to measure with conventional sampling technology, but are thought to be important 'hot spots' of enhanced grazing activity on phytoplankton and elevated predation rates by copepod predators. My research focused on analyzing data from a Zooglider mission conducted in three distinct regions off the eastern coast of Spain. During the 39 day mission, the glider executed 352 dives during which 395k images were collected. After the cruise, this data was processed into 1.39M labels of individual plankters labeled by a machine learning algorithm with concurrent CTD measurements. Access to this metadata was provided via a PostgreSQL database. To facilitate this analysis, I utilized pgAdmin for preliminary queries and managing the extensive dataset generated by the Zooglider. For backend processing, I employed Python and SQLAlchemy, while Plotly and Dash were used to create interactive visualizations.
Ultimately, I deployed three web applications to the lab’s web server for data presentation and analysis. My first web application summarizes the cumulative amount of data collected and was used to validate the database contents. My second web application allows for targeted browsing of the collected images, and was used to verify that the ML algorithm's labels were reasonable. My third web application plotted vertical profiles of CTD data and ML label counts, allowing me to determine the presence/absence, location, depth, and thickness of the layers being analyzed. Results indicated the presence of Copepod Thin Layers across all regions. In the majority of the vertical profiles, a bimodal distribution of the No. Copepods / L were detected, which had not been resolved before through other zooplankton sampling methods such as towed plankton nets. By capturing one image every 10 cm, the Zooglider enabled the high-resolution identification of Copepod Thin Layers in the ocean. I also found that environmental factors such as chlorophyll-a concentration, marine snow, and density gradients were found to correlate with these layers.
2023
Matteo Biscotti

The method of extraction used for characterizing dissolved organic matter in seawater has been shown to have great implications on the final characterization of a sample. Because of this, I utilized the more contemporary and widely used PPL solid phase extraction for the Scripps Pier samples, as well as the now less widely used C₁₈ solid phase extraction. Additionally, I used both acidified and unacidified PPL solid phase extraction but only acidified C₁₈ solid phase extraction. My goal was to isolate compounds that are more efficiently extracted by a given solid phase extraction method as well as compounds that were found only under certain environmental conditions. Following the extraction, I used liquid chromatography and coupled mass spectrometry before decoupling the mass spectral data using MZmine 3. Finally, the data was matched against the GNPS database, and I used Sirius for compound class annotation.
While the results were largely as expected, with a more diverse set of compounds being extracted by PPL filters, this is good as I had shown that the more contemporary method is generally more efficient. Few trends emerged correlating to environmental conditions, but this was expected due to the limited amount of variation in the data being analyzed. Still, the wide variety of compounds annotated represented only a small number of the total compounds present in the samples, further indicating that changing the methodology will yield different characterization. I am now doing additional work to characterize samples from stations along California’s coastline.
Elizabeth Burstein

Understanding the plankton community in the California Current is important for answering questions about the success of fish larvae as well as other members of the ecosystem.
I investigated potential food sources for anchovy larvae in the California Current. Anchovies and other fish larvae select prey to genus or species level, but most studies of ocean-caught larvae only identify prey to class or order level. Using the NOAA CalCOFI Oceanographic Genomics dataset, generated through metabarcoding, I found correlations between anchovy larvae and individual plankton strains.
Using R to calculate Spearman correlations, I found taxa that appear with anchovy larvae. Some of these were previously known food sources, e.g. Dinophyceae dinoflagellates, especially Gymnodinium and Lepidodinium. Clausocalanus furcatus copepods also correlated with the larvae, and their nauplius stages may be important food for the larvae as they grow older. Many other species were detected, but not identified because their sequences were not present in the reference dataset. There is lots of work to be done identifying these taxa, as well as looking at fish larval stomachs to confirm causal relationships with the potential food taxa.
Meghan Kaschner

Meghan Kaschner investigated prokaryotic and eukaryotic community structure and diversity in the California Current Ecosystem (CCE) over a seven-year period, from 2014 to 2020. Three main questions were asked. First, how does the structure of prokaryotic and eukaryotic communities change in the CCE over time? She suspected, after literature review, to find community structure variation between cooler periods (2018 to 2020) and warm periods (there was a warm anomaly and El Niño from 2014 to 2015). Second, how does diversity scale with total biomass, and is this pattern consistent between prokaryotes and eukaryotes? Broadly, she expected diversity to increase as total biomass increased. Last, how does the diversity of prokaryotic and eukaryotic communities change over time? She hypothesized that over time, diversity is stable and change in diversity is correlated between prokaryotes and eukaryotes. She used Amplicon Sequence Variant (ASV) data collected by the quarterly California Cooperative Oceanic Fisheries Investigation (CalCOFI) cruises, as part of the NOAA-CalCOFI Ocean Genomics (NCOG) Project. ASVs provide taxonomic information and absolute abundance at higher resolution than previous standards (i.e Operational Taxonomic Units); each ASV is an exact DNA sequence.
In reference to the first question, she determined relative abundance for each taxonomic order per cruise using the absolute abundances provided with each ASV. Paired with a mean chlorophyll time series—greater abundance of reads was correlated with higher chlorophyll concentration in both prokaryotic and eukaryotic communities—she assessed relative changes in biomass over the seven-year period. To test the second hypothesis, she investigated ASV richness, Shannon diversity index, and ASV evenness. Each metric was plotted against the total number of reads per cruise. To address the third question, she created line plots of diversity metrics over the seven-year period.
She observed a shift in prokaryotic and eukaryotic community structure where dominant prokaryotic ASVs switched between warm and cooler periods, and the dominant eukaryotic order increased in magnitude during the cooler period. She found no relationship between diversity and prokaryotic biomass, but as eukaryotic total biomass increased, richness, diversity, and evenness decreased. Last, she found that diversity changed over time for both prokaryotes and eukaryotes (more variable), and only richness first differences were correlated between the two communities.
Next, she intends to rarify the ASV data, look at community composition overturn with respect to time with the Bray-Curtis index, and conduct spatial analyses by dividing the samples into four known bioregions. Adding 12s ASV data to incorporate fish species diversity is also of interest.
Carlo Makeever
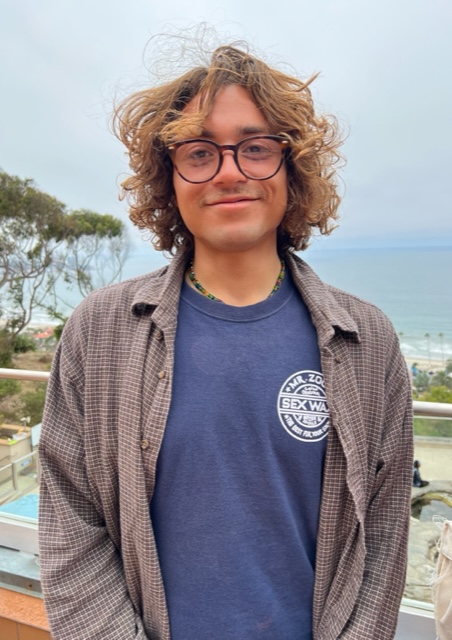

For my REU summer project I worked in Dr. Mark Ohman’s lab investigating spatial changes in mesozooplankton grazing. This summer, I had the opportunity to work with Dr. Art Miller through the REU program, focusing mainly on the ecological responses to changing environmental conditions within the California Current Ecosystem (CCE). Utilizing a numerical model, we examined how the CCE responds to various environmental shifts, such as El Niño events and reduced upwelling, just to name a few. My tasks ranged from learning Matlab for data processing to conducting statistical analyses on model outputs, which included variables like temperature, nutrient levels, and various types of plankton. Our key findings revealed what the general behavior is of the CCE throughout the year. For instance, we observed that diatoms thrived during the upwelling season when nutrients were high, and zooplankton growth was most significant when diatoms were present. These insights help us understand how the CCE might respond to future environmental changes, particularly in the context of climate change. I am incredibly grateful to Dr. Art Miller, Colleen Petrik, Anthony Wilson, and the entire CCE mentorship program for their invaluable guidance. This REU experience has not only equipped me with essential skills in data analysis and scientific inquiry but has also inspired me to explore further research opportunities in marine ecology.
Angeles Rios

This summer, I worked in the Scripps Acoustic Ecology Lab under the mentorship of PhD student Michaela Alksne, where I investigated Baird’s beaked whale presence throughout the North Pacific using passive acoustic data. Baird’s beaked whales forage at depth, making it difficult to rely on visual observations alone to determine their spatio-temporal distribution. However, this species, like all toothed whales, produces a unique echolocation click that can be used to identify them near underwater recording sites. Using MATLAB software built in the Scripps Whale Acoustics Lab, I was able to identify and label Baird’s beaked whale echolocation clicks from this acoustic data by analyzing spectra, which compares the frequency and received level of a sound. I created a time series for sites in the California Current Ecosystem, the Olympic Coast National Marine Sanctuary, and the Gulf of Alaska to display Baird’s beaked whale presence. Most notably, my results suggest a seasonal pattern in Baird’s beaked whale presence off Point Conception, as the majority of detections only occurred from spring to early fall. Additionally, there was a greater relative presence in the Gulf of Alaska when compared to the other regions, pointing to a potential species hotspot to be further investigated. This work will contribute to researcher and policymakers’ understanding of Baird's beaked whales' spatio-temporal distribution and, once built upon, may yield further insights into their population dynamics and broader ecological role.
This REU program allowed me to explore research for the first time and gain hands-on experience with data analysis that will serve me in my next steps. I would like to thank Michaela Alksne for being an incredibly supportive mentor and for helping me gain confidence in my work. Thank you to my PI Dr. Simone Baumann-Pickering and the Scripps Acoustic Ecology Lab for allowing me to be a part of such a collaborative and friendly environment this summer.
Jack Saade

Over the course of the summer, I worked on the development and optimization of a new prototype ion selective electrode (ISE) and its housing to accurately measure ocean pH levels. I used a variety of programs to accomplish this goal. Labview was used to read raw data and translate it into meaningful values; Matlab, to organize the data into coherent visuals and calculate constants; and Solidworks, to continually design and 3D print a pressure sealed housing for the ISE.
This was all done with the goal of improving on existing technology. Through iterative design and a mix of chemical, electrical, and mechanical engineering, the proper chemistry of the ISE was found, the minimum number of components were used, and the optimal housing dimensions were calculated. The next steps going forward are to optimize the electronics to eliminate noise and manufacture an official housing.
Iris Smryl

This summer, as an REU intern, I worked with Art Miller on physical-ecological models\ of marine microorganism populations off the West Coast. The ocean model I was working with was provided by Colleen Petrik, and was forced by observed atmospheric data from the years 1958-1991. Having no prior coding experience, I began to learn Matlab from Grad student Anthony Wilson as a high-level computer language process. From there, I learned to read the data files of the models into Python and interpret them. Then I began to plot snapshots of the data at specific times, and make movies or GIFs of the output. The farthest I got this summer was to begin looking at statistics of the model output, like seasonal cycles of monthly means, and also looking at population anomalies in the data for each month of a year.
These models are useful for a number of things, with the main two being aggregating a lot of the large amounts of physical-ecological variables present in nature, down to ones relevant to concisely addressing a given question, and for understanding ecosystem responses to the physical processes that drive the changes. In our case, we can narrow the variables down to the ones that control nutrient flow, like POC, temperature, and depth, or the density of the diatoms
that the zooplankton graze on. Now, we can look at relationships between ecology and specific abiotic factors, like seasonal upwelling, El Niño/La Niña, and long-term physical climate changes associated with global warming. This model also allows us to look at biotic relationships like competition between the diazotrophs and diatoms, or lead-lag relationships between the diatoms and zooplankton.
I would like to thank the CCE LTER program, and everyone involved with the Miller lab for giving me such an amazing opportunity and experience over the summer, and for the support and guidance they provided. I plan to continue the research I started in this program into the school year and beyond, using the skills and connections I made in this program. I would also like to give a huge thank you to Art Miller as well for being such an amazing mentor, teaching me so much about a field I began to enjoy, and for the amazing support as well! I can’t wait to continue working with him!
2022
Hanna Budroe

For my REU this summer, I worked in the Décima Lab under the guidance of graduate student Dante Capone, using DNA metabarcoding to analyze patterns of zooplankton diversity and community composition here in the California Current Ecosystem. Molecular methods such as metabarcoding present a faster and potentially more accurate alternative to morphological characterization, which allow us to identify zooplankton species from high volumes of complex samples. For my project this summer, I worked with metabarcoding data for both mtCOI and 18s rRNA "barcodes," performing various ecological analyses in the language R. Within R, I was able to identify community clusters in zooplankton correlated with both a cross-shore and size-fraction gradient, characterize the community composition within these clusters, and also compare the performance of the two different barcodes across various taxa.
Establishing a baseline survey of zooplankton diversity and community structure is important, as these current patterns were largely dictated by changes in temperature and nutrients, and these conditions may change as climate change progresses. Given that various zooplankton taxa differentially contribute to biogeochemical processes, such as carbon sequestration, understanding these community patterns are essential for predicting and understanding ecosystem functioning.
I am incredibly grateful to the CCE-LTER REU program for giving me this opportunity to immerse myself in the world of research, and to the Décima lab for their tireless support this summer. A special thanks to Dante Capone, who not only patiently mentored and coached me through all the frustrating code, but also brought me homemade bread and iced me countless times. All these amazing experiences this summer have inspired me to pursue graduate school, and given me the confidence and skills to follow these passions.
Maria Durazo

This summer I had the opportunity to work in the Aluwihare lab, under the mentorship of PhD student Ralph Torres, to study the diel variability of marine metabolites. Organic matter in the marine environment is distributed into particulate organic matter and dissolved organic matter. For this project we focused on the analysis of dissolved organic matter (DOM), which is composed of different types of important compounds varying in size, such as proteins, lipids, amino acids, etc. It is important to note the large contribution that the microbial community has to the DOM pool, this helps us understand the variability of compounds at different time points as photosynthesis and respiration changes occur throughout the day.
This summer, we performed a week-long study off the Ellen Browning Scripps Memorial Pier, collecting seawater samples at 12am, 6am, 12pm and 8pm. From these samples we were able to collect nutrients and dissolved organic carbon, as well as use a solid phase extraction method to be able to retain and analyze the organic compounds. Using a Liquid Chromatography- Mass Spectrometry method (LC-MS), we obtained the mass spectrum of several compounds that were later uploaded to a database to be able to identify the metabolites. Only 85 out of 2,685 metabolites were identified, but using other identification tools we were able to predict the potential compound classes. A variety of compounds in our samples displayed diel variability and were found to be important molecules for biochemical processes. Not only were we able to find metabolites, but also found the presence of anthropogenic compounds, which can help us think about the impact of human activities on the appearance of these compounds. Using the identification of metabolites and information about the chemical environment, collected through nutrient and dissolved organic carbon concentrations, it is possible to hypothesize the importance and function of these compounds in the marine environment.
I would like to thank Dr. Lihini Aluwihare, graduate student Ralph Torres, and the Aluwihare lab for their guidance and support this summer. Coming into this internship I had little knowledge about oceanographic sciences, but throughout the summer they taught me how to use all the tools I needed to know for this project and introduced me to another very interesting area of research. I am very grateful to have had the opportunity to participate in the CCE-LTER program as an REU. I have learned more about the process of research and the elaboration of a scientific project that has shown me many skills that I will continue to use in the future.
Kassadi Holloway

For my REU summer project I worked in Dr. Mark Ohman’s lab investigating spatial changes in mesozooplankton grazing. Along our California coast we have upwelling, which is when cool, nutrient-dense water from deeper depths rises, and a filament is when this water travels out into the sea like a river, before eventually mixing with the open ocean. From satellite images we can see that phytoplankton bloom along the filament from all the nutrients. What I researched was how their predators, the zooplankton, react to such changes. My overarching question was: how does grazing pressure change along a coastal upwelling filament?
In August of 2019, cruise P1908 followed the Pt. Sur Filament using drift trackers, conducting day and night plankton tows. These mesozooplankton were then run through different sized filters to place them in 5 unique size fractions, from 5000 to 500μm. In the lab, I removed the samples from liquid nitrogen, cut them into a specific fraction, and placed them under the microscope for sorting. I carefully removed any detritus, such as algae, to ensure only the plankton remained, taking note of dominant or unusual species along the way. Next I placed the clean sample in acetone, sonicated it to break up the tissues, and placed it in the freezer for an hour, to allow any gut pigments to release into solution. After centrifugation, I removed the supernatant and ran this through a fluorometer. This machine uses the specific fluorescence of chlorophyll to relay how much of the pigment was in the sample; these values are used to calculate the grazing rate of the plankton. I found that the mesozooplankton have a higher
mass-specific grazing rate inshore, however the phytoplankton here are so well-nourished from the filament they can offset this predation. Therefore, the grazing impact is actually highest offshore. I also identified that the smaller size fractions, <1000μm, dominate grazing inshore, but pyrosomes, which are large colonial plankton, control grazing offshore. I’m incredibly grateful for the opportunities this program gave me. Not only did I learn many useful lab and data analysis skills (and plankton species, of course), I also saw what it takes to create and pursue a research question. This experience has helped me explore paths for my future, and for that I want to thank everyone in the CCE-LTER program that helped to make this happen, and especially my mentor Mark Ohman, as I will be staying on staff in the fall to continue this research.
Zoie Jones

As a CCE LTER undergraduate intern, I spent my summer working in the Scripps Acoustic Ecology Lab with postdoc Annebelle Kok and graduate students Ella Kim and Gabby Arrieta, where I studied whale calls and fish choruses in La Jolla Canyon. My primary goal was to assess baleen whale and fish vocal behavior in the canyon by analyzing data collected via a Zooglider, a modified spray glider featuring a hydrophone (i.e. underwater microphone) that moved throughout the canyon. I used programs in MATLAB to view the data collected in the form of Long Term Spectral Averages or LTSAs, which look at sound over time and allowed me to identify what whale and fish species are present in the canyon. I then created visualizations in R to assess the diel periodicity of the calls and choruses produced, and where fish are in the water column.
I am incredibly thankful for being able to be a part of the CCE LTER REU program as I was able to work with scientists of various disciplines, analyze and interpret data first-hand, and learn about the many ways marine systems can be studied! Thank you to my mentors Annebelle Kok, Ella Kim, and Gabby Arrieta who helped me tremendously when working on this project, the Scripps Acoustic Ecology Lab Principle Investigator Simone Baumann-Pickering for her insight, Mark Ohman and Sven Gastauer who collected the data I analyzed, and the Scripps Acoustic Ecology Lab, Scripps Institution of Oceanography, and CCE LTER program for giving me the tools and opportunity to work as an REU this summer.
Katarina Kaminsky

Over the summer, I had the opportunity to work in Dr. Mark Ohman’s lab under the guidance of PhD student Stephanie Matthews. I spent half of my summer in the lab and the other half performing data analysis utilizing DNA metabarcoding to assess biodiversity and abundance of zooplankton in the North Pacific Basin. I worked with DNA extracted from samples taken at different depths using a vertical multinet from across the North Pacific. The samples used in this analysis were taken from the epipelagic, upper mesopelagic and deep mesopelagic regions of the water column. To determine the identity of these samples, the CO1 mitochondrial DNA gene and the 16S ribosomal DNA gene were amplified using PCR, gel electrophoresis and purification techniques. After the samples were sequenced, the R software program was utilized to perform community similarity analyses in various depth zones and at different latitudes. After the data were processed and visualized, a latitudinal biodiversity gradient was identified in which biodiversity tended to decrease as we moved north away from the equator. The data were also a subset, and a preliminary analysis was done on the distribution of the mollusc community in the sample locations. Spatial patterns within the mollusc community were observed, however, further analysis is required to draw broader conclusions from this data.
I feel incredibly grateful for this opportunity which has provided me with the skill set and confidence to execute research, laboratory work and data analyses. The community and knowledge that was cultivated during my summer as an REU is invaluable to me as I continue in my academic career and beyond. I want to specially thank my mentor, Stephanie Matthews, who made my experience so enriching and fun, as well as all of CCE LTER for giving me this opportunity and the people at SIO and JCVI for all of the support along the way!
Kailey Ramsing

As part of the CCE LTER REU program, I had the opportunity to work in the Palenik Lab studying cyanobacteria in the San Diego Bay and off the Scripps Pier. Cyanobacteria are important microbial primary producers in the ocean, and I specifically studied the genus Synechoccocus, which is one of the most abundant oceanic genera of cyanobacteria. I collected samples from the SIO Pier in order to continue the study of microbial population dynamics that the Palenik Lab has been conducting for almost two decades. I also ran multiple experiments using Synechoccocus isolates from the San Diego Bay in order to understand function in the unique bay environment. First, I used a heat block to understand the preferred temperatures of each strain and to study if the isolates can tolerate the warm bay environment. I also ran a darkness experiment to understand the dark tolerance and recovery of Synechoccocus, as the San Diego Bay experiences a high amount of mixing and turbidity that can trap microbes with little to no sunlight for periods of time. We found that all isolates could survive at least 10 days in the darkness and recover fully. We plan to continue this research to understand the limit of darkness that the isolates can tolerate to connect to dynamics in the San Diego Bay
I am very grateful for this opportunity and would like to thank the CCE LTER program and the Palenik Lab for allowing me to continue research this summer, as well as all members of the Palenik Lab for their continuous support. This experience has provided me with valuable skills and experience in marine microbiology that I am confident I will continue to use in the future.
Kaitlyn Reardon

This summer with the CCE LTER and Franks Lab, I was able to utilize MATLAB to look into important physical oceanographic factors, especially relating to upwelling and the upwelling season, along the California Current System. Using geostrophic velocity satellite data, I asked two main questions: What are the regions along the California coast where, on average, flow during the upwelling season is directed offshore, and what are the temporal patterns (when/how often) in offshore flow probability at these locations?
Initially, I looked at the monthly probability of offshore flow using geostrophic velocities along 34-43°N. To determine which values were considered offshore flow, dependent on the geography of the coast, offshore flow had to be at an angle within a 90-degree threshold with the center value being perpendicular to the coast. Visually, there were key ‘hotspot’ points along the west coast where it seemed there was a higher probability. This initial observation helped me segway into temporal patterns; by looking at individual capes and points along the coast, I was able to identify certain patterns, such as seasonal patterns, in some locations. Most notably, northern latitudes, such as those by Cape Blanco, had lower offshore probability in the winter months in comparison to the summer months. In the mid-latitudes, such as those by Point Año Nuevo, there was more variability across all seasons, but looking within the ten-year time scale, there were higher probabilities in offshore flow in some years versus others. Finally, looking at the lower latitudes, such as near Point Conception, there is an annual trend of higher probability of offshore flow in the spring and summer months each year.
In regard to spatial patterns, I used Hovmoller plots to highlight if certain latitudes had higher overall probabilities, and how those changed or were similar in comparison to other years. After analyzing each year, it was noted that because the California Current System is impacted by climate oscillations and decadal scales, such as ENSO, NPGO, PGO, and ONI, and the upwelling season, spatial patterns vary immensely, and therefore, predicting offshore flow varies at each location yearly.
I would also like to thank Shailja Gangrade, my mentor this summer, as well as my PI Peter Franks and the Franks Lab for such an enjoyable experience this summer and all the help I received from the lab group on a weekly basis. It was awesome being able to apply the information and skills I’ve learned in the Oceanic and Atmospheric Sciences major here at SIO to something so important in the real world, and the REU program and Shailja were amazing steps in making this happen!
2021
Victoria Burns

As part of the REU, I had the opportunity to work in the Décima Lab developing a workflow for assessing pteropod shell dissolution. Pteropods are an ecologically important type of planktonic gastropod, whose beautiful, fragile shells are made of aragonite, the more soluble form of calcium carbonate. Because of this, pteropod shells are acutely prone to dissolution in waters undersaturated with respect to calcium carbonate, such as those present in the CCE during periods of upwelling or those projected as a direct consequence of ocean acidification. Since pteropods are among the most susceptible to changing carbonate chemistry, they act as an important bioindicator, serving as a bridge between the biological impact of OA and the chemical reality of it.
My project consisted of investigating current methods in assessing pteropod shell state and adapting them for implementation here at SIO. It was an iterative process of exploration and refinement: each stage, from sample preparation to the final image analysis, had a number of considerations that needed to be assessed. I was taught how to use the SEM by Dr. Greg Rouse, who generously shared of his time, resources and expertise in this arena. In order to measure the affected area, the SEM images were analyzed using software coded for the purpose. I developed a preliminary image analysis workflow to quantify the variable nature of dissolution. Future work would apply these objective measures of dissolution to pteropods collected from a range of OA conditions. This experience was absolutely incredible due, in large part, to my amazing mentor, Dr. Moira Décima, and the wonderful community here at Scripps, including the Rouse Lab and Linsey Sala and crew at the PIC.
Anissa Garcia

During this past summer, I worked alongside Darcy Taniguchi’s lab at California State University San Marcos. We performed full diel dilution experiments off the Ellen Browning Scripps Memorial Pier in La Jolla, CA to study the growth and grazing mortality rates of marine plankton. I was also given the opportunity to partake in a month-long research cruise through the California Current Ecosystem Long Term Ecological Research (CCE LTER) program to continue performing full dilution experiments in the Pacific Ocean. The full dilution experiments were accomplished by incubating seawater at different dilution factors (whole seawater and filtered seawater ratios) and analyzing samples during a day and night cycle (12- and 24-hour periods). Seawater incubated at different dilution factors influenced the encounter between heterotrophic protists (grazers) and phytoplankton (prey). Collecting samples after 12 and 24 hours helped us estimate the grazing mortality and growth rates of the entire community over time. Other samples were also taken to characterize the environment and planktonic community.
Grazing mortality and growth rates, based on chlorophyll values, were higher during the day compared to the night for all regions sampled. The net growth rates were higher during the night in the open ocean, in contrast to the nearshore sampling location which had higher rates during the day. Further analysis of auxiliary samples will help distinguish which communities were present at each sampling site, thus allowing us to investigate what phytoplankton and heterotrophic protists were more prominent during the day versus the night.
I would like to thank the National Science Foundation for funding me this summer. I would also like to thank CCE LTER, Scripps Institution of Oceanography, and UC San Diego for their support. Thank you to Sydney Plummer for being a tremendous graduate student mentor and for helping me while on the research cruise. Thank you also to the crew on the R/V Roger Revelle for helping the science party collect samples, the Stukel, Diaz and Décima labs, Natalie Yingling, Dante Capone, Anya Stajner, and Monica Thukral for providing help during the research cruise, Robin Westlake Storey for facilitating the REU summer program, and Darcy Taniguchi for welcoming me into her lab in 2019; it is because of Darcy I love helping in research!
Chiara Hayes

This summer I worked in Dr. Brian Palenik’s Lab. The focus of my project was to analyze the growth rate of Synechococcus species at different temperatures, using isolates from San Diego Bay and the California Current. In addition to this I also learned and utilized a pathway and genome database known as BIOCYc (https://biocyc.org/) to determine the presence of genes and chemical reactions amongst the Synechococcus species being analyzed. The goal of this aspect of the project was to determine potential differences amongst the various species to better understand how they might adapt and grow under different thermal conditions. Understanding the abundance and growth rate of Synechococcus bacteria is very important as they are important contributors to major nutrient cycles in the ocean such as the carbon cycle and are major producers of oxygen. To conduct growth experiments, I set up and measured the fluorescence of eighteen tubes of a specific Synechococcus isolate over a duration of two weeks. The tubes were placed in a temperature gradient ranging from fifteen degrees Celsius to thirty degrees Celsius. After all of the fluorescence readings were recorded for a given species, I developed a model in Excel to encapsulate Synechococcus growth vs. temperature to visually understand the best thermal circumstances for Synechococcus growth. In addition to this, I would analyze genes and reactions that various species had in common using BIOCYc.
This project has better helped us to understand the temperature ranges for different Synechococcus species. It has also helped to determine similarities in adaptations amongst species in the San Diego Bay area and around the Scripps pier. Not all of the gene similarities have yet been found, but many have been predicted to share genes that allow them to undergo adaptations under various conditions of temperature. Being affiliated with the CCE-LTER program was a very valuable experience for me. With it I have learned more about the diversity and majority of adaptations that Synechococcus species can undergo. I have partaken in a vast majority of laboratory skills that have enhanced my knowledge of important protocols. I have also gathered researching skills to interpret gene sequences and reactions of organisms in large genome databases such as BIOCYc.
Anna Mothersole

This summer I worked in the Scripps Acoustic Ecology Lab under the guidance of graduate student Michaela Alksne, and I had the opportunity to research species specific trends in dolphin presence and absence while trying to answer the question of if stock-specific echolocation clicks truly correspond to two genetically distinct Pacific White-Sided dolphin populations - a northern population and a southern population. I was also able to investigate trends in Risso’s dolphin presence and absence at a recording site close to the channel islands marine sanctuary.
Previous studies have shown that there are two different echolocation click types produced by Pacific White-Sided dolphins in the Southern California Bight, so Michaela had trained a neural network to identify those two click types (LoA and LoB), as well as Risso’s dolphin echolocation clicks, among other marine sounds. Passive acoustic monitoring was used to collect the raw data, and I then annotated the clicks labelled by the neural network to ensure an accurate dataset. Pacific White-Sided dolphins were primarily studied at a site in the Southern California Bight (though data from along the U.S. west coast was available), while Risso’s dolphins were studied at a site further north near the CCE2 mooring. Using generalized additive models in R, we were able to use environmental data from the CCE2 mooring to model Risso’s dolphin presence and absence compared to temperature, salinity, julian day, and lunar phase. Our results suggest that the Pacific White-Sided dolphin LoA click type corresponds to the northern population of the dolphins, as the click type is found all along the west coast of the United States, while the LoB click type was only found at the southernmost recording site, suggesting that it corresponds to the southern population. Also, the models of Risso’s dolphin data show their presence and absence to be significantly correlated with salinity, julian day, and lunar phase. While these observations and models of Pacific White-Sided and Risso’s dolphin presence and absence trends can give some insight into the dolphin’s population distribution and habitat use, further research is needed to understand their underlying causes.
I would like to give special thanks to Michaela Alksne and everyone in the Scripps Acoustic Ecology Lab for all of their help, and for allowing me to take part in research with them. I have not only learned about skills and tools useful for marine acoustics research, but I was also able to gain a lot of coding and modeling experience, particularly in R, and I’m excited to be able to take these skills with me into my future scientific career.
Sandy Nguyenphuoc

As an REU student this past summer, I had the opportunity to work in the Burton Lab under the guidance of graduate student Michaela Labare. I utilized DNA barcoding to monitor California ichthyoplankton across five locations, where I was able to assess the biodiversity and abundance of pelagic spawning fish. To accurately identify these fishes, I amplified the CO1 mitochondrial DNA gene and the 16S ribosomal DNA gene with PCR and gel electrophoresis techniques. The data we collected was then compared to previous data collected from these sites (2019 and 2020) in order to elucidate patterns in diversity and egg abundance spatially (latitude) and temporally. Once compared, we found that Scripps Pier has consistently had the highest egg abundance and species diversity. We also found that the Northern anchovy, a commercially important species, has a spawning season that is consistent with previous years. Furthermore, our data shows that locations north of Point Conception tend to have lower levels of diversity compared to more southern locations (Choi et al. 2021).
I am incredibly grateful to have been able to participate in this REU, as it has provided me with first-hand experience of working in a laboratory setting and analyzing my own data. Aside from strengthening my technical skills, I am also thankful for the support system that I have found this summer. Overall, this REU has shown me that there is still much to learn and that I have only grazed the surface of this field. Thank you to Ron Burton, Michaela Labare, Natalie Faivre, and the Burton Lab for giving me such a valuable opportunity this summer.
Charlene Ruiz

This summer I had the opportunity to work for Dr. Andrew Barton's lab to research changes in the phenology of harmful algal blooms (HABs) at Scripps Pier. The project is dependent on data from SCCOOS' harmful algal bloom dataset (2012-present) and W.E. Allen's historic dataset (1919-1939), both of which provide weekly cell counts of phytoplankton measured at Scripps Pier. Studying changes in phytoplankton bloom phenology provides further insight on the variability of changes impacting the productive Southern California Bight from events such as climate change. Comparing and contrasting the two datasets allowed me to see how bloom phenology might have transformed within the past century, especially in the timing of its peak abundance and seasonal peaks. I used R to perform data analysis, in which I located and identified algal blooms in both datasets based on a particular threshold. This summer I looked primarily at four bloom-forming dinoflagellates shared between the two datasets, to observe how similarities or differences in the rates that these dinoflagellate taxa may have changed across time. Data visualizations were furthermore created via R to identify changes in bloom timing and magnitude alongside dynamic environmental conditions, such as temperature and salinity.
Working as a California Current Ecosystem REU intern has given me the opportunity to see what it was like to dedicate myself to research full-time, as well as explore the extent of my research interests. I am grateful to Andrew Barton and the Barton Lab for their guidance and mentorship, and I look forward to further developing the skills I've gained over the summer.
Bryant Tran

This past summer, I worked in the Semmens Lab under the mentorship of Dr. Erin Satterthwaite, PhD student Tammy Russell, and Dr. Brice Semmens. I attempted to find a causal link between marine animal stranding data and phytoplankton counts in the California Current Ecosystem, plus its surrounding regions. Information on phytoplankton composition was taken quarterly, 4 times each year, and originated from CalCOFI (stranding data was also consolidated to have total seasonal counts). Hundreds of species were documented, but the research focused on the pennate diatom Pseudo-nitzschia, variants of which are associated with harmful algal blooms. The data were confined to parts of the ocean from Point Conception down to San Diego, and ranged from 2013 to 2018. Using data analysis techniques, R, and Python, I plotted autocorrelation functions, correlation maps, and linear models of the data. From this, I determined that there was a positive correlation between stranding counts of the California sea lion and Pseudo-nitzschia populations, with a time lag of around one year. These results showed a distinct relationship with seasonality, with strong peaks during the spring and summer. With further research, I hope to create a model to help predict when harmful algal blooms occur, and determine the environmental drivers for them.
Taking part in the CCE-LTER internship was incredibly rewarding, and I would like to thank Brice Semmens for recommending me into the program. All of my mentors are amazing people, and I hope to make everyone who was a part of this proud with the work I do in the future as I pursue a graduate degree.
Marley Weiss

Over the summer, my research at Scripps was focused on measuring trace levels of total dissolvable iron (TdFe) in seawater samples. These samples were taken from the P1908 CCE LTER process cruise, collected from various depths along the path of an upwelling filament near Pt. Sur. These seasonal upwelling currents bring deep waters rich with nutrients such as iron to the surface, which are often limiting agents in the rate of primary production but can also limit primary producer growth altogether. Samples were acidified to pH ~1.8 to dissolve potentially bioavailable iron off the surfaces of particulate matter and tested for TdFe concentration using flow injection analysis. This data was then processed using MATLAB and formatted into Ocean Data View, a software designed to visualize section plots in the marine environment. The data gathered during the REU internship provided insight on the seasonal variations of iron levels in the open ocean and in near-shore, high productivity regions. By comparing these values for iron concentrations against those related to primary productivity, such as particulate organic carbon (POC) and biogenic silica (bSi), it becomes possible to understand the biological links between these elemental cycles in the ocean. This aligns with the goal of CCE LTER research by measuring iron levels over the course of many years to formulate a comprehensive understanding of the biogeochemistry of the region.
I would like to thank Katherine Barbeau for all her help throughout the program and Kiefer Forsch for mentoring me on the processes of trace metal analysis. The experience I gathered from this internship was invaluable, giving me the opportunity to understand the responsibilities of independent researchers as well as collaborating with other researchers in a professional environment. My practices with flow injection analysis, MATLAB, Excel, Ocean Data View, and analytical chemistry are important skills that I can apply towards the future of my career. This internship has helped me on my journey through higher education and may possibly lead to a future in oceanographic research.
2020
Alicja “Ala” Najduch
This past summer, I worked in Dr. Andrew Allen’s Lab under the guidance of PhD student Rob Lampe. I analyzed phytoplankton diversity and community composition in the California Current Ecosystem using data collected from three sample sites offshore San Diego between 2009 and 2011. These samples were collected before, during, and after phytoplankton blooms, including one associated with the red tide-forming dinoflagellate Lingulodinium polyedra. Using 18S ribosomal rDNA sequencing, I characterized the phytoplankton community composition from the different sites and across time. As predicted, the sites were predominantly either dinoflagellate or diatom dominated. Using transcriptomics, I further characterized the relative abundance of taxonomic groups based on their mRNA abundances. To provide further environmental context for our data, I also analyzed satellite imagery of sea surface temperature and chlorophyll-a concentrations in the region. Collectively, these data will provide insights into the responses of different phytoplankton taxa during bloom events in the region.
I would like to thank Rob Lampe and the Allen Lab for this research opportunity and guidance this summer. Working as a CCE-LTER intern was an incredibly valuable experience. I gained a better understanding of the applications of genomics and transcriptomics, the different species of local phytoplankton, and the marine environment. Additionally, I gained a lot of practical experience manipulating data in R, and greater access to the amazing academic network at Scripps. Working in the lab has helped me solidify my plans to attend graduate school, and I hope to continue working on this project with the Allen Lab.
Iris Kubler-Dudgeon
 This summer as an REU student, I had the opportunity to work in Dr. Amina Schartup’s lab developing a model of mercury accumulation in phytoplankton. Mercury is a neurotoxin that accumulates in marine food-webs. Long-term exposure can cause chronic health impacts on general cognitive functioning. The largest increase in concentration of mercury between trophic levels occurs between seawater and phytoplankton. The mechanisms driving this increase are still poorly understood, particularly in the Pacific Ocean. To better understand the transfer of mercury in the water column I developed a model that could account for changes in seawater temperature, nutrient and light availability, and depth, allowing us to see mercury accumulation in phytoplankton over time as the population changes. These environmental changes most impact the lifetime of mercury in the oceans and its accumulation by phytoplankton. Using this model with projected future sea water temperatures could help predict mercury transfer into higher trophic levels and assist in managing health standards as climate changes. Throughout this research experience I have improved my coding skills and learned about physical and biological variables affecting plankton population dynamics and mercury distribution in the oceans. I gained insight into the process of building and adapting a mathematical model which I will take with me to future research.
This summer as an REU student, I had the opportunity to work in Dr. Amina Schartup’s lab developing a model of mercury accumulation in phytoplankton. Mercury is a neurotoxin that accumulates in marine food-webs. Long-term exposure can cause chronic health impacts on general cognitive functioning. The largest increase in concentration of mercury between trophic levels occurs between seawater and phytoplankton. The mechanisms driving this increase are still poorly understood, particularly in the Pacific Ocean. To better understand the transfer of mercury in the water column I developed a model that could account for changes in seawater temperature, nutrient and light availability, and depth, allowing us to see mercury accumulation in phytoplankton over time as the population changes. These environmental changes most impact the lifetime of mercury in the oceans and its accumulation by phytoplankton. Using this model with projected future sea water temperatures could help predict mercury transfer into higher trophic levels and assist in managing health standards as climate changes. Throughout this research experience I have improved my coding skills and learned about physical and biological variables affecting plankton population dynamics and mercury distribution in the oceans. I gained insight into the process of building and adapting a mathematical model which I will take with me to future research.
Juliet James

This summer I worked on a project in the Aluwihare lab, the focus of which was determining if the difference between modeled and measured carbon to chlorophyll a ratios could be used as a proxy to separate phytoplankton carbon concentrations and detrital carbon concentrations within the directly measured bulk suspended particulate organic carbon (POC) concentration. The modeled values we used are from a study by Qian Li et. al in which they created a model to predict only the phytoplankton-associated carbon concentration in a water parcel based on light and nutrient data. This phytoplankton-associated carbon can also be referred to as living carbon. The measured values include both the living carbon, as well as all sources of detrital carbon such as dead organic matter, runoff, etc. Thus, we wanted to know if the total measured POC minus the living carbon yields a reasonably, oceanographically consistent value for detrital carbon concentrations across the Californian Current Ecosystem LTER domain (2006-2017).
We looked at variation in our predicted detrital carbon values throughout the water column by region and saw high variability in its concentration in nearshore waters, but very low and invariable concentration offshore. Upon comparison to total measured carbon profiles, we found that, in general, this comparison of carbon to chlorophyll ratios does seem to provide a good estimate of detrital carbon concentration throughout the water column. The next step is to find a measurement that can independently verify detrital carbon concentration so that we can compare this to our estimated values to determine how accurate our estimates are.
Nicolas "Nico" Concha-Saiz

This summer I worked in the Choy Lab under the guidance of postdoctoral researcher Elizabeth Hetherington. My project focused on examining differences in the elemental composition (nitrogen, carbon, carbon to nitrogen ratios, and moisture percentages) of micronekton (organisms 2-20 cm) in Monterey Bay, CA. I used previously collected data to compare elemental compositions of five phyla (Arthropoda, Chordata, Cnidaria, Ctenophora, Mollusca), and I compared variability within and between taxa. Throughout the summer, I conducted literature reviews on micronekton community composition, elemental analysis, and learned the statistical programming language R. In R, I learned how to make figures and analyze data using linear models. I found significant differences between hard and soft-bodied taxa, where gelatinous animals (e.g., cnidarians, ctenophores) had a higher moisture content and lower carbon and nitrogen than more muscular animals (e.g., fishes, squids). Examining trends in elemental composition and moisture content can tell us more about the biology and feeding ecology of animals that make up food webs in the pelagic ocean. We can also use this data to evaluate differences in elemental composition between active and passive predators, and animals that occupy different depth habitats. This research will be useful for future estimates of biomass and carbon flow through the deep pelagic ocean. Being part of the CCE-LTER was an incredible and valuable experience. I learned a lot about food webs and micronekton in the CCE. This summer project provided me with research experience and has inspired me to continue to conduct research in the field of marine biology. I want to thank Anela Choy, Elizabeth Hetherington, and everyone who stepped up to the challenge of a virtual REU.
2019
Garret Schmid

I spent my summer as a CCE-LTER REU intern working within the Dr. Martz Chemical Oceanography Lab. I am about to finish my senior year as a Marine Science student at Cal Poly SLO and found the research experience very valuable to my career. I worked on the Smartfin project, which integrates sensors within a surfboard fin for oceanographic data collection in the coastal ocean. Working closely with Dr. Phil Bresnahan, my objective for the 11-week summer was to test multiple pH sensors to understand their capacity to be incorporated within a small surfboard fin. While the project is niche (integrating ISFET pH sensors within a Smartfin), understanding the characteristics of specific pH sensors has far ranging applications to better assist carbonate monitoring and to explain the variability of ocean acidification in the marine environment. Overall, I learned basic electrical engineering skills, wrote software to communicate with microcontrollers, designed 3D printed jigs and housings, ran controlled experiments to test the pH sensors, analyzed pH voltage output, and built a few functioning Smartfins which involved soldering circuit boards and potting fins with fiberglass and epoxy resin. Not only did I gain a vast range of skills, but I was able to build a network of connections at Scripps, both within and outside my direct lab, that will be foundational to my personal and professional career as I am applying to graduate programs.
Lawrence “Rence” Balitaan and Yunchun “Pauline” Pan


This summer, we worked with Dr. Art Miller, under the mentorship of PhD candidate Nathali Cordero-Quirós, to better understand the biophysical interactions of the California Current System (CCS). For our internship, we utilized MATLAB to plot and analyze 11 variables from the ROMS-NEMURO model: temperature, salinity, zonal and meridional current velocities, nitrate and silicate concentrations, three types of zooplankton grazers (categorized by size), and splitting up chlorophyll into two parts—small and large phytoplankton.
We first analyzed data at a single location off the coast of Baja California Sur. We plotted single depth profiles, depth averages, climatologies, anomalies, and standard deviations of the aforementioned variables. We then repeated this analysis at two separate box locations at the northern and southern portion of the CCS. Once we accomplished this, we started creating more visually appealing maps of the entire CCS. We took the anomalies from our model and correlated them with the Oceanic Nino Index, a leading indicator scientists use to determine if the sea surface temperature at the equatorial Pacific Ocean experiences warming or cooling conditions. We analyzed each variable at various lag times to characterize any relationships between our model with the observational data. In these correlation maps, we specifically observed how a development or decay of one variable affects another. Lastly, we created composite plots of each variable at specific and mean depths during El Niño and La Niña conditions. Using these composite plots, we analyzed each parameter both individually and collectively to discover and characterize trends.
Pauline: As an Applied Math undergraduate, I practiced my programming skills by extracting and manipulating data stored in 4D matrices and learned different MATLAB plotting tools throughout this internship. Moreover, I learned about the different physical and biological variables and understood the process of upwelling and its effect on biological variables. This background knowledge helped me analyze the plots I made.
Rence: As a student of the new undergraduate Oceanic and Atmospheric major at UCSD, I contributed my extensive knowledge on biological, chemical, geological, and physical oceanography. Moreover, I improved my limited coding skills. I learned how to manipulate large data sets in MATLAB that allowed me to analyze the data. Through this internship, I developed more insight on how all the contiguous parts of the California Current System interact and respond to the effects of climate change.
Natalie Faivre

This summer I worked in Ron Burton’s lab under the guidance of current Masters Student Emma Choi. This ongoing project monitors fish spawning at the marine protected area (MPA) off the Scripps Pier and was started by a Masters Student in 2012 to investigate the effectiveness of this particular MPA. In addition, data collected from an MPA can help researchers understand the effects of climate change on marine life. For the project I looked at fish eggs to observe spawning patterns and draw conclusions on how climate change might be affecting species diversity and population numbers. Recently, this project was expanded and we are now looking at fish eggs and environmental data from 5 additional sites (Santa Cruz, Santa Barbara, Cal Poly-SLO Avila Pier, Newport Beach, and Santa Monica) to be able to make further comparisons.
To get the data, we used a plankton tow to collect the fish eggs, performed DNA extraction, and then a polymerase chain reaction to obtain the amplicons that will be sequenced using BLAST sequencing database. Once we know the number of fish and what the species are, we can do data analyses to begin to hypothesize explanations for any patterns we notice.
Working as a CCE intern was an incredibly valuable experience. I would like to thank Ron Burton and everyone in the lab for being so supportive and helpful, and for giving me a place to apply my knowledge in a tangible way. My experience has inspired me to pursue this type of research in the future, and I am incredibly grateful for my time spent here this summer.
Tristin Rammel

This summer I had the opportunity to work in Brian Palenik’s lab, concerned with phytoplankton dynamics and ecology. I was involved in analyzing the twice weekly, 20+ years-long time series of microbial genetic diversity and abundance off the Scripps Pier. Due to the size of the dataset, I focused on cryptophytes, a ubiquitous eukaryotic taxa with a role in primary production and trophic transfer of energy. Using DNA extraction methods and bioinformatic software, I characterized the dominant clades of cryptophytes off the Pier and within the San Diego Bay. I also analyzed cryptophyte cell densities over three years using flow cytometry, in order to divide relationships between cryptophytes and other relevant microbial taxa in CCE waters. Participating in this research taught me a wealth of molecular and bioinformatic methods with which to investigate microbial communities, as well as practical field sampling methods that I will be able to take with me in my future research career.
2018
Emma Choi

This summer I worked in Dr. Ron Burton’s lab researching fish spawning trends in the La Jolla Marine Protected Areas. The project was initiated in 2012 and serves to assess the efficacy of Marine Protected Areas and identify general trends in fish spawning behavior. I joined the project under the guidance of Masters Student, Elena Duke, to continue identifying fish species spawning near the Scripps Pier. This consisted of collecting fish eggs, amplifying their DNA by PCR, and sending the DNA out to be sequenced. Once sequences were obtained, they were matched to a species using GenBank. This molecular method of identifying fish eggs complements data regarding local fish species obtained by diver surveys. It is a useful way to gather information about the ecosystem in the La Jolla Marine Protected Areas because diver surveys account for adult fish, but do not contribute to information on spawning behavior.
Working as a California Current Ecosystem REU intern has given me the opportunity to obtain hands on research and collaborate with experienced scientists in the field. I would like to thank the Burton lab for providing me with a welcoming community and educational learning environment.
Fernando Gil

This summer I had the opportunity, and pleasure, to work for the Martz Lab in MESOM at SIO. The core work performed by this group includes data analysis of the CalCOFI Cruises and bottle samples. The CalCOFI takes data measurements every four months along the coast of California. Our lab, which collaborates with them, is then able to process that data and hopefully make conclusions from them. Our main motivation for this research and work is the persistence of ocean acidification. Ocean acidification, also known as "the other CO2 problem," is the continuing decrease of the pH (becoming more acidic) of the Earth’s oceans. This is caused by the large and increasing amount of CO2 absorbed by the ocean which then shifts the pH toward a more neutral position, rather than basic. This negatively impacts the ocean's ecosystem as many fish and coral rely on a balance of carbonate ions in order to create their shells and skeletons. For this reason we look to utilize this data to create prediction models and bring awareness, in hopes that we can combat this issue before it is too late. I was assigned the task of creating a program with MATLAB that would perform the Quality Control aspect of data processing. This included creating several live scripts that would perform multiple aspects of the quality control process. Utilizing the system’s built-in gas standard system, which runs a known gas standard in order to check the accuracy of the analyzer when analyzing the SeaWater pCO2, I was able to create a three point regression in order to correct the underway data. Following corrections made about assumptions that were utilized throughout the collection of data, I was able to calculate the alkalinity of the samples, which could then be used to calculate the pH. Although the Underway system does collect a pH value, this measurement is obtained through arbitrary constants and therefore needs to be adjusted. I also included a Gantt chart script, which plots a Gantt chart in order to show which components of this system were functioning at any given time along the length of the cruise.
I am very grateful for this opportunity. Working at the Scripps Institute of Oceanography has always been this magical place, filled with people like me who are passionate about the ocean, and this summer showed me just how right I was.
Sara Ramirez

This summer as an REU intern, I had the opportunity to work with Dr. Art Miller and PhD candidate, Nathali Cordero-Quiros. I was able to do research on the California market squid to look at possible ways to project species capacity. The market squid is very important both commercially and ecologically in the California Current Ecosystem, as understanding the physical-biological interactions that affect species abundance are crucial for future fishery management and ecosystem stability. To do this, I began to look at habitat suitability modeling and applied this concept to spawning suitability. Because market squid are restricted by their environment during spawning the model was constructed using the species-environment relationships of bathymetry, substrate affinity, and temperature; ranking spawning zones on how suitable it is based on those criteria.
This becomes important as El Niño/La Niña produce different ranges of area that are suitable for spawning (due to changes in ocean temperature, among other changes), leading to fluctuations in species abundance. Using this model and projecting future temperature changes could potentially assist with how fisheries are managed.
This research project has been very valuable in launching my scientific curiosity, and I would like to thank the Miller Lab for giving me the opportunity to explore this topic independently. I gained a lot from this summer experience and I am thankful to have been a part of the Summer 2018 cohort.
Niv Anidjar

As an REU student in the Barbeau Lab, I worked under the mentorship of PhD student Kiefer Forsch. As part of his goal of constraining a value for biological iron demand in the CCE, I used a fluid injection analysis machine to measure dissolved iron concentration in seawater samples taken during process cruise P1706. I used these concentrations along with ADCP velocity data in order to determine values for cross-shore advective flux of dissolved iron. These measurements are also useful for understanding changes in distribution of primary producers, as iron is episodically the limiting nutrient in the California Current Ecosystem, which can restrict growth of plankton.
While working in this lab, I learned the procedures for preventing contamination of trace metal seawater samples, how to interpret biogeochemical oceanographic data, and how to form hypotheses based on this data by examining relevant published literature. This program was an irreplaceable experience for me. The opportunities it provided for me to engage with current oceanographic research have helped me to better understand my academic interests, as well as to be more comfortable with the idea of going into graduate school in this field in the future.
Kellie Lemoine

During this past summer, I worked under the guidance of PhD student, Lauren Manck, in the Barbeau lab at Scripps Institution of Oceanography. The focus of my summer research was to understand the role of siderophores in microbial iron acquisition. Siderophores are compounds that strongly bind and transport iron in microorganisms.
I ran growth experiments with a siderophore synthase knockout mutant strain of Alteromonas bacteria and a normal wild type strain of Alteromonas bacteria that still retained the siderophore synthase gene. I grew the two bacterial strains in marine broth and then transferred to replete pc+ media before iron-limiting the cells. I then grew the bacteria in media with different iron compounds and measured the cell density every three hours in order to plot the growth curves. The results suggest that in the California Current Ecosystem, where upwelling may be a source of sedimentary iron, siderophores may play a significant role in helping bacteria uptake inorganic iron.
I would like to thank Lauren Manck for teaching me so many valuable molecular biology tools and techniques this summer. This internship experience has reinforced my passion for scientific research and has spiked my interest in oceanography as well. I look forward to implementing the skills I learned this summer in my future scientific work.
2017
Cristina Curiel

This summer I worked under the mentorship of Dr. Art Miller in plotting and analyzing model data output by the Darwin model. This model combines physical and biological aspects of the CCE modeled by real data in the 2000s to better understand the California Current System and its impact on biology. The Darwin model is an emergent model different from models commonly used in biology. Rather than inputting a single phytoplankton or zooplankton for the model to output a result, the Darwin model uses 20 different groups of phytoplankton and 8 different groups of zooplankton and outputs results based on the conditions of the time-based model given by a series of equations. The phytoplankton and zooplankton groups are grouped based on size, growth factors and nutrients, so therefore depending on the conditions, some phytoplankton may be more successful than others.
This project focused on physical structures in the ocean known as eddies, with interest in comparing cross-frontal model data to what is seen in the field. For example, if one group of phytoplankton is successful within an eddy of the model data, do we see the same happening in similar conditions in the field? In order to find the answer to whether or not eddy transect data in the model match real data, it is important to take the model data and synthesize it into visual contour plots. This was done through MATLAB, a computer programming language I was completely unfamiliar with before this program. After learning my way around MATLAB, I was able to make contour plots that presented model data of the ocean surface for a given month and day, for any variable in the model data such as phytoplankton, temperature, salinity, and various nutrients. This created a visual to understand the relationship between physical conditions and biology. I am really grateful for this opportunity as it really expanded both my experience and my knowledge of ocean science. I got to experience science and research first hand as well as learn more about research at Scripps Institution of Oceanography.
Gabriela Haddad

This summer as an REU intern, I had the opportunity to work in Dr. Ron Burton's lab with master's student Elena Duke. Our project involved studying the spawning patterns of fish in our local Marine Protected Area, particularly in two habitats- the kelp forest and off the Scripps pier. In order to do this, we collected fish egg samples from these habitats and molecularly identified them by extracting their DNA, amplifying it via PCR, and sending that DNA off for sequencing. We then analyzed those sequences using databases such as NCBI and identified each egg on the species level. From these data, we were able to obtain results showing species compositions in the kelp forest and off the Scripps pier, which species were spawning in which habitats, when their spawning periods were, and how environmental effects such as El Niño affected these fish populations. We were also able to analyze these results for multiple years and thus draw long term conclusions from the multi-year data.
Over this summer, I was able to learn valuable research and interpersonal skills and gain further awareness about our California Current Ecosystem. This experience allowed me to achieve a deeper understanding of the scientific world and its endeavors to better understand and care for our oceans. I also had the opportunity to work with and learn from outstanding individuals who no doubt have helped me along these steps in my scientific career. I would like to thank the Burton Lab for allowing me to have this invaluable experience and Elena Duke for being an outstanding mentor.
Aysell Medina

This summer, I was given the opportunity to work as an REU at Scripps Institution of Oceanography with Dr. Lihini Aluwihare and PhD candidate, Sara Rivera. During these few weeks, we studied the heterotrophic bacteria in the California Current Ecosystem using samples taken from the CCE-P1706 cruise. Our goal was to measure the bacterial abundance and compare the size of the bacteria to previous years.
To study the bacteria, I took the fixed samples and used a Blue Angel to filter the bacteria and mounted them onto microscope slides stained with Vectorshield containing DAPI that stained the bacterial DNA. I then imaged the samples using a C1 Nikon light microscope and sized the bacteria using the NIS-Elements program. The exported data from the program was then normalized, and I counted, measured, and calculated the carbon content for each cell. Although our results are still being analyzed, we are also considering running our samples using flow cytometry, observing the influence of grazers within the system, or observing the bacterial strains present in our samples for validation and further understanding of the current data we possess.
This experience helped me understand how scientific observations can be approached and carried out into projects. I also learned how many individuals with a diverse background and areas of study can all complement and equally contribute toward advancing our knowledge of the California Current Ecosystem. Most importantly, my REU summer experience reinforced my desire to further my education and continue in the research field. I would like to thank Sara Rivera, Dr. Lihini Aluwihare, and the CCE LTER REU program for offering me such a rewarding and valuable opportunity.
Kaytlyn Taglialavore

During the summer of 2017 I had the pleasure of working as an REU at Scripps Institution of Oceanography under Dr. Lihini Aluwihare and PhD student Brandon Stephens. The focus of my study was on colored and fluorescent dissolved organic matter in the California Current Ecosystem. The data I worked with were from the previous cruise in June, CCE-P1706. In order to analyze the data I ran all 225 samples collected throughout the cruise (4 cycles, 3 transects) in a spectrophotometer for absorbance. This helped gain information on chemical composition and molecular size of colored dissolved organic matter. Then, the samples were run in a fluorometer to collect the excitation and emission outputs for each sample. These samples were then coded through MATLAB using a process called PARAFAC. The MATLAB PARAFAC then output components of the FDOM (fluorescent dissolved organic matter). This information was then analyzed with reference to different information regarding the samples like distance offshore, DOC concentrations, depth, etc. This data was then put into terms using a quinine sulfate standard and can be analyzed with prior cruise information on FDOM (from CCE-P1604, P1408, Student cruise 1507) to gain information about the chemical transformations that happen with reference to different circumstances (like “Blob” year or El Niño).
This summer internship experience was incredible and helped my ability to understand how the ocean is connected to current climate circumstances. It helped me grow a respect for research and the ability to analyze complicated processes that are influenced by thousands of possibilities. It helped reinforce my passion in sustainability and understanding the effects that humans have on the environment, and specifically the ocean.
Maggie Wang

During this summer, I worked in Dr. Brian Palenik’s Lab with the guidance of PhD student Maitreyi Nagarkar. I analyzed Synechococcus diversity in the California Current Ecosystem using filters from the four cycles observed in the P1604 cruise. After extracting DNA from the filters, I used primers to amplify the ITS region and sent it to RTL for sequencing. I used a program to assign taxonomy to the sequences and discovered that the offshore sites had greater amounts of Synechococcus from clade IV, and the coastal site had more clade I sequence data. In addition, there wasn’t much of a difference in Synechococcus diversity when I compared the surface sample data to deeper depth data from the same site.
I would like to thank Dr. Brian Palenik and Maitreyi for this research opportunity I had this summer. I gained a better understanding of the different species of Synechococcus and their environment, as well as a closer look into the approach of research. I plan to continue working on this in the Palenik Lab.
2016
Bretton Coppedge

This summer in the CCE LTER REU program I worked in Dr. Katherine Barbeau’s lab under the mentorship of PhD student Lauren Manck. The Barbeau lab is primarily interested in studying trace metals in the ocean and in particular, iron. For that reason, we studied iron uptake in a very important marine bacterium, Alteromonas macleodii, which previous work has suggested is responsible for a significant part of iron cycling in the CCE. We created a gene similarity network that helped us identify which genes Alteromonas most likely uses to take up iron and then we analyzed the expression of these genes through qPCR analysis, under iron limited conditions.
During my time in the lab I learned many valuable techniques such as how to prepare slides for epifluorescence microscopy, how to culture bacteria and do growth experiments with them, and I also learned how to do PCR/qPCR and gel electrophoresis. This program was also helpful when it came to getting advice from graduate students about the application process of applying to graduate school. Both the techniques I learned and the advice I was given will be invaluable in my career moving forward.
Elena Duke

As an REU student in the Burton Lab at the Scripps Institution of Oceanography, I studied the spawning activity of fish populations in two of San Diego’s Marine Protected Areas (MPA) using DNA barcoding with PhD candidate, Alice Harada. We collected fish eggs from the end of the SIO pier using a 505 micron net. The fish eggs were sorted and counted and tored in ethanol at four degrees Celsius. We crushed the eggs and put them in AE buffer to extract the DNA which we then amplified with PCR using species specific primers and COI gene primers, and later 16S primers. We sent the PCR product to be sequenced and compared the data to reference sequences in GenBank NIH. This study shows that DNA barcoding is a good alternative to other fish population studies. These data are able to show the seasonal changes in spawning activity and changes in community composition over time.
This summer I learned a lot about DNA barcoding and the molecular tools scientists’ use as well as ecological knowledge of the fish populations that are in two of San Diego’s MPA’s. I’m thankful for the opportunity to work with the California Current Ecosystem Project and to work in Ron Burton’s Laboratory. I plan to continue working in Ron Burton’s lab on this project.
Wiley Wolfe

My name is Wiley Wolfe and I am a senior at Oregon State University majoring in chemical engineering with a minor in oceanography. During the CCE LTER REU internship, I worked in the Martz lab focusing on ocean chemistry. The goal of my summer was to determine the chemical spatiotemporal variability of Agua Hedionda Lagoon. This means that I was looking to see how much the chemistry of the water in the lagoon changed with space and time. To do this I used a WavepHOx sensor which, every 15 seconds, measures pH, dissolved oxygen, temperature and salinity. The WavepHOx was paired with a GPS watch so the time and location of each measurement was recorded. The sensor is designed to be mounted on the bottom of a paddle board. Therefore, a large piece of my summer was spent on a paddle board, making for a pretty great summer internship.
After the data was collected, I analyzed it using MATLAB. The data collected showed that there was significant variation in the lagoon. This was especially evident in one corner of the lagoon that was a salt marsh, a type of wetland. We believe that most of this variability is due to the daily cycles of respiration and photosynthesis. This variability is important for things such as accurately determining the carbon flux or productivity in the lagoon. Some future work I would like to do on this project is, first - calculate productivity using an assumed alkalinity. Second, further analyze the calibration data for the WavepHOx. Finally, take more measurements in the lagoon using a sensor that is able to measure two components of the carbonate system to completely constrain it. Overall my time in San Diego was a ton of fun and great experience, I learned a lot about the carbonate system, electronics trouble shooting, computer programming, and field data collection.
India Dove

During the summer of 2016, I worked in Dr. Mark Ohman’s Lab under the guidance of graduate student Jennifer Brandon on a microplastics project. Specifically, my REU project involved researching the multi-decadal changes in microplastic sedimentation rates in the Santa Barbara Basin (SBB). The question was whether or not microplastic sedimentation rates changed through a long-term timescale and if there was a distinct time horizon when plastic first emerged in the sediments. The SBB presents a unique opportunity to analyze microplastic sedimentation through time because it’s anoxic at the basin bottom and consequently no biological organisms mix the varved sediment layers that correspond to distinct years in time. I analyzed a sediment core extracted from the SBB which was chronologically dated from 2009-1834. The core was previously processed into 83 vials and each vial corresponded to a year or two in time. After enumerating, imaging, measuring, and recording various characteristics about each of the 1,128 microplastic pieces extracted under the microscope from vials 1-13, corresponding to years 2009-1992, it was apparent that fibers were the dominant particle type throughout each vial. Interestingly, the fibers were of an array of colors (blues, teal, tan, copper, pink, and green), however white was the principal fiber color. The majority of the microplastic pieces extracted from the sediment samples of 2009-1992 ranged in size from 0.5-1.0mm. From the data obtained thus far, a potential connection may be seen between microplastic sedimentation and annual rainfall in the Santa Barbara area; sediment vials corresponding to years 2005 and 1997-99 had a greater number of microplastic particles compared to other vials and years 2005 and 1997-98 were distinctly heavy rainfall years for the area. However, further analysis of the remaining sediment vials needs to be completed in order to fully understand the effects of contamination and accurately calculate the sedimentation rates of microplastics through time in the SBB.
I would like to thank Dr. Ohman and Jenni for the invaluable research internship I had the privilege of experiencing this summer. I gained a further understanding of the process-oriented approach of research and a greater appreciation of the importance of the California Current Ecosystem. Additionally, my awareness was expanded regarding the extent at which microplastics infiltrate and proliferate throughout the ocean and consequently the ramifications they pose for marine life. The implications of the microplastic research greatly aligned with my sincere passion for marine conservation. This summer has reinforced my future career aspirations to use all the means at my disposal in order to mitigate anthropogenic threats jeopardizing the integrity of the ocean and endangering life within its realm.
Brooke Rasina

During the summer I had the honor of working as an REU at Scripps Institution of Oceanography under the guidance of PI Dr. Lihini Aluwihare and PhD student Sara Rivera. My focus of study was on the heterotrophic bacteria in the California Current Ecosystem. The question we attempted to answer was whether El Niño had an effect on the bacterial abundances and bacterial carbon content. The bacterial abundance and carbon data from P1604 (El Niño year) was compared to similar data from P0605 (normal year), P0704 (normal year) and P1408 (“Blob” year). In order to analyze the collected samples I warmed them, filtered the samples, stained them with Vectorshield containing DAPI and mounted them on slides. Next I imaged 20 fields of view per sample on a Nikon-C1 microscope and utilized the images to count and size the heterotrophic bacteria. The collected count data was then converted into bacterial abundance per milliliter, and the collected size data was converted into carbon content using pre-determined relationships between size and carbon content. The aforementioned data was then compared to the past cruises to determine if El Niño may have had a noticeable effect on the bacterial abundance and carbon content in the microbial loop. Our initial findings suggest that most of our data fit in the range of observations from P0605, but significant spatial differences were observed that still need to be explored in the context of other available datasets to discern any possible El Niño impact.
This summer was truly an incredible experience and it helped reinforce my desire to continue on in my higher education and pursue a PhD. I am going to continue on in Dr. Lihini Aluwihare’s lab through an independent study to complete the data set and further analysis. Dr. Aluwihare, Sara Rivera and I also want to compare our results to different data collected about the microbial loop so we can achieve a more complete picture of what is happening and why certain trends are present. I look forward to seeing this project through to completion and getting a more well-rounded idea of the research process.
Casey Graff

During my REU with the CCE-LTER project over the summer, I worked in the Ohman lab at SIO studying machine learning techniques for the automatic classification of digital zooplankton images. Classification and quantification of plankton samples is necessary to understand plankton distributions, create oceanographic models, and study population ecology. Automating this process can save significant time and resources. Under the guidance of Dr. Mark Ohman and PhD candidate Jeffrey Ellen, I wrote computer algorithms that were able to surpass previous accuracy benchmarks. The zooplankton images that I worked with were collected and preserved as part of the CCE-LTER project and digitized using a high-resolution scanner, called a Zooscan. Presently, images collected using the Zooscan are sorted using an older, less accurate algorithm and the results are verified and corrected manually, a resource intensive process. Improving the accuracy of the algorithm has the potential to greatly reduce the amount of time invested in the manual correction step.
Furthermore, future image collection techniques may necessitate accurate, fully-automatic classification due to the greater volume of images that may be collected. One example of this is the submersible Zooglider, a prototype, which takes thousands of images while traveling through the water. My improvements over existing algorithms were obtained by the use of a Convolutional Neural Network (CNN) which incorporated geometric features and environmental metadata about the collection of the images. This geometric and environmental information was previously not used in the CNN and was shown to improve the overall classification accuracy when added. This is a step towards the goal of having a completely autonomous classification process.
This experience has helped me to improve my understanding and implementation of the scientific process. During my REU, I gained a new appreciation for the interdisciplinary aspect of science. I interacted with many individuals of highly-focused and varied backgrounds who work closely together to advance our understanding of the oceans. I also was able to improve my approach to problems by applying a strict scientific mindset; developing a testable hypothesis followed by executing well-designed experiments to validate or invalidate the hypothesis. This was the first opportunity that I've had to pursue a scientific question full-time, a distinct difference from regular coursework. Given the substantially greater amount of time to explore my hypothesis I was able to investigate many more of the related questions that are raised during the investigative process. I thoroughly enjoyed my time working in the Ohman lab this summer and it has provided me a unique insight into how research is conducted.
Jennifer Beatty

This summer, I had the privilege of working in Dr. Michael Landry’s lab under the supervision of PhD student Ali Freibott. My project was estimating the ciliate biomass in the California Current during the “Blob” and El Niño warm water anomalies. Ciliates are heterotrophic plankton that represent up to one third of the heterotrophic biomass in the California Current. They are extremely fragile and require a different filtering process than other plankton. I used samples that were preserved using acid Lugol’s gathered during the CCE process cruises of 2014 (“Blob” year) and 2016 (El Niño year). I filtered the water onto slides and analyzed them using light microscopy methods. Then, following previously outlined methods, I counted ciliate cells and calculated the carbon biomass. We found that the ciliate biomass during the El Niño cruise was higher than the biomass found during the “Blob” cruise. There was a wider range of biomass in samples from El Niño than in samples from the “Blob.” Additionally, we found that the water temperatures in the “Blob” cruise were generally higher than the El Niño cruise, and that chlorophyll levels had a comparable range between the respective cruises.
2015
Linda Tong

As an REU student in the Ohman Lab at Scripps Institution of Oceanography this summer, I studied mesozooplankton grazing in the California Current Ecosystem under the guidance of Dr. Mark Ohman and graduate student Jennifer Brandon. I analyzed zooplankton samples from process cruise P1408, and then examined possible factors affecting spatial variability in mesozooplankton grazing and sought to determine an appropriate metric for approximating the factor of food concentration.
In the laboratory setting, I handled frozen zooplankton collected during summer of 2014 and analyzed the samples for gut fluorescence on fluorometers—the measurements of chlorophyll-a and degraded gut pigment served as an indication of zooplankton grazing through ingested phytoplankton. After working through the five cycles of cruise P1408, I used the data to compare possible factors affecting patterns of zooplankton grazing. Several variables were tested, including distance from shore; size of zooplankton; and concentration of phytoplankton available for consumption. The same was then done with preexisting data from cruise P1208. Focusing on the factor of food concentration, I analyzed data collected directly from cruises P1408 and P1208 to find a metric of phytoplankton availability that could represent a clear relationship with zooplankton grazing rate.
My participation in the CCE LTER REU program has allowed me to gain invaluable experience and knowledge. I have developed a greater appreciation for the California Current Ecosystem, and am now better informed of topics varying from local zooplankton species to graduate school programs. I am so thankful to have had the opportunity to learn so much in one summer, and to have worked with amazing people at such a beautiful location!
Raul Martinez

As my first formal internship, this summer I had the honor to work and learn in Kathy Barbeau’s lab at Scripps Institution of Oceanography, performing trace metal analysis under the guidance of third-year PhD student Angel Ruacho. Copper is a trace metal of concern in San Diego Bay and coastal waters off Southern California because it is toxic for phytoplankton species at high concentrations. The main input of copper comes from antifouling boat paint, which is used to prevent marine organisms from growing on boat hulls. Coastal runoff is another significant source. I had the opportunity to explore San Diego Bay on a small boat to collect samples all over the bay at 7 different locations, as well as sampling from the Scripps Pier, a coastal monitoring station. Our studies included the characterization of copper-binding organic ligands by using electrochemical measurements in order to determine possible indicators of copper toxicity. The method used is called Competitive Ligand Exchange – Adsorptive Cathodic Stripping Voltammetry (CLE-ACSV). From my experience, I gained an understanding of the basic application of this method from a chemistry perspective and also how the data are analyzed to determine copper and ligand concentrations, in addition to binding strengths of ligands.
This exposure to research with oceanographers helped me to understand the importance of all disciplines in science. As an engineering major, I found this experience not very convincing at first, but ultimately it was truly an astonishing opportunity to remind myself how all science is interrelated and that every scientist has something to learn from another. I never realized all of the research that is being done on our oceans and the importance that it has for our planet. After this summer, I definitely felt more inspired to appreciate science and its power to answer the many challenging questions that currently surround us.
Maria Winters

During my CCE-LTER REU experience this summer I worked with Dr. Art Miller on a physical oceanography modeling project. I analyzed output from a combined biological-physical model of the California Current Ecosystem called the Darwin Model. This model represents the changing populations of phytoplankton and zooplankton in the CCE over time, as well as changing physical variables such as temperature, ocean velocity, and salinity. By utilizing output from this model, we aimed to identify correlations between ocean physics and phytoplankton communities in this ecosystem within physical ocean structures called eddies. More specifically, we hoped to identify why different phytoplankton communities grow differently at different depths within an ocean eddy and to see if emergent biological features in the model exhibit similar cross-frontal transitions as observed in the CCE. Specifically, I utilized the computer programming language MATLAB to plot output from the model in the form of Network Common Data Form (NetCDF) files. Future work needed on this project will be to use statistical analysis to quantify the model output and then compare it to observed data from the CCE. The CCE is one of the most productive ecosystems in the world, and by better understanding and improving models to represent it, we can better predict the effects of short-term weather events and long-term climate change on the biological populations within it.
I truly enjoyed working on this project because it combined biology, physics, and computer programming- all important aspects of the degree in environmental engineering that I am pursuing at UC San Diego. I had the opportunity to greatly improve my MATLAB skills, but I also learned about modeling, oceanography, and climate science research being conducted at SIO. Through the CCE-LTER REU program, we received tours of different collections and laboratories at SIO, and we also got the chance to meet a wide array of scientists, faculty, and graduate students, which allowed me to learn about research in fields other than my own. Overall, this experience greatly expanded my knowledge about the research fields I am interested in, and I plan on continuing to pursue my passion in preserving and protecting the environment utilizing what I learned during my experience at Scripps Institution of Oceanography.
Vivian Han

During my three months with the CCE LTER internship at Scripps Institution of Oceanography, I had the opportunity to work in the Palenik Lab under the guidance of PhD student, Maitreyi Nagarkar, and PI - Dr. Brian Palenik. My research focused on two questions during the three month period. The first one focused on the population dynamics and the clade distribution of Synechoccocus, an ecologically important cyanobacteria, at the SIO pier. To approach this question, I collected sea water samples at the SIO pier regularly, extracted total environmental DNA, and amplified the RpoC gene, which is useful for distinguishing between the various clades of Synechococcus. I created clone libraries of RpoC, which allowed me to analyze the clade distribution during Synechoccocus population blooms and crashes at various time points.
My second research question investigated whether Gymnodinium, a genus of photosynthetic dinoflagellates that has been shown to exhibit mixotrophy in other studies, grazes on Synechoccocus. I set up co-culture experiments to create nutrient-deprived conditions in order to induce grazing. The result, measured through fluorometry and microscopy, suggested that in vitamin deficient conditions, Gymnodinium seems to have a possible grazing relationship with Clade I of Synechoccocus.
My three months as a CCE LTER REU was a very inspiring experience. It opened the gate of oceanography research and informed me of different fields of research related to oceanography. It also better prepared me for graduate school and led me to understand what research will be like as a future career.
Abdalrahman Alrayyes

This summer was a great step towards pursuing my future endeavors, where I had the honor of working under Dr. Ralf Goericke on the CalCOFI group’s oceanic research. I was learning something new every day, whether it was the chemical dynamics of the ocean, or just a new way to visualize different relationships using code. My job function was a data analyst, and I was essentially going through decades worth of pH data to either confirm that the data were true, or to decipher new relationships that can allow us to better understand the ocean’s chemistry.
For the first half of my internship, I was examining data from 2 pH sensors used on CalCOFI cruises, and comparing them to a set of theoretical pH values. I was able to find that both the sensors were faulty for multiple reasons, some of which were physical limitations and calibration issues. When that was discovered, I was back at square one, where what I had was only theoretical. Like any scientist in that situation, I started making sense of what was available to me, seeing how factors such as nitrate and oxygen contents are related to pH. Through that, and with the help of members of the CalCOFI group, who I owe many thanks to, I was able to develop a relationship that calculates the surface water pH of the ocean based on 3 factors. This will come in great aid for scientists trying to study things like ocean acidification or calcification.
As a chemical engineer, the computer and software skills that I learned to develop models, examine sensor reliability, and regress relationships for over 60 years worth of data, were invaluable experiences that I am very grateful for. Hence, I would recommend anyone trying to get into the scientific field to pursue a position in the CCE-LTER REU program.
2014
Taylor Wirth

I had the opportunity to work in the Todd Martz Lab under mentor and Ph.D. candidate Phil Bresnahan. The lab develops autonomous ocean sensors, most notably the SeaFET pH sensor and SeapHOx pH & oxygen sensor, and is currently developing new sensors for total alkalinity and dissolved inorganic carbon. The need for autonomous, high precision ocean chemical sensors is growing as the oceanic uptake of carbon dioxide increases, thus decreasing ocean pH by 0.002 pH a year. This small change can only be measured with very precise sensors, such as the ones the lab develops. These sensors have been distributed to collaborators in SIO and all over the globe. A new area of study is the very near-shore environment, where Phil has implemented a method to measure near-shore pH by attaching a SeapHOx sensor to a stand-up paddleboard (SUP).
My project was to take the existing SeapHOx and modify it to be lighter, smaller and more hydrodynamic. To do this, I used computer aided design (CAD SolidWorks) to design a new housing that was less than half the size and weight of the previous version. It also includes a dome that allows the sensor to move through the water with minimal drag. The new design has a half-cylinder shape, allowing it to attach to potentially any flat surface. Along with the new design, a new battery pack was implemented to allow for recharging, which was not possible with the previous versions. The modifications highly simplify the deployment and storing process of the sensors. The sensors are now currently being built in bulk numbers to be sold to collaborators who are in need of one, such as the Air Sea Interaction Research Lab who will attach one to a Wave Glider. The new design is called the WavepHOx.
I also participated in data collection and data mapping while on a SUP with a SeapHOx. I determined the best method to mapping the data and portraying the spatiotemporal differences in sea surface pH (up to 0.2 pH change in a single day!). The WavepHOx has huge potential for many applications, such as collecting baseline measurements to compare anomalous events to (rain, high surf, algae blooms...), long distance/open ocean autonomous sensing, and local outreach programs for people interested in surfing and maintaining the ocean's chemistry. The CCE-LTER REU program gave me invaluable experience in ocean sensor development here at SIO. I am very grateful for the opportunity and all that I have learned while participating in the REU program.
Maya Land

This summer, I worked in Mike Landry’s Lab with mentor Moira Décima. My work consisted of learning a relatively new method of secondary production estimation and implementing it on the CCE LTER research cruise in August (P1408) to measure the chitobiase-based production rates of crustacean zooplankton communities. This method is based on the fact that crustaceans release a molting enzyme (chitobiase) into the water column when the molt. Assuming that chitobiase is microbially degraded at a certain decay rate, the rate of chitobiase production should equal the slope of the decay curve. Thus, chitobiase levels in the sea water along with the rate of decay calculation can provide a productivity estimate. The decay curve was achieved by sampling seawater, incubating it for 24 hours and measuring chitobiase following a protocol developed by Sastri and Dower (2006, 2009). Further analysis will determine bioproductivity rates using chitobiase production estimates and crustacean zooplankton biomass (or total chitin measurements).
The CCE LTER REU experience has been unforgettable and a truly eye-opening experience. I was able to come away not only with valuable lab experience, but also with a greater understanding of graduate school, biological oceanography as a field, and future research career prospects as well.
Kari Ann Yoshida

As part of the CCE LTER REU internship this past summer, I had the opportunity to work in the Palenik Lab at the Scripps Institution of Oceanography, under the mentorship of Dr. Brian Palenik and grad student Maitreyi Nagarkar. As well as helping with time series pier collections and other lab duties, I conducted guided research on the survival of Synechococcus at differing ocean depths. From the collection and filtration of sea water samples to the organization of specific Synechococcus DNA sequences, throughout my REU experience I was able to gain a greater perspective on the methods and techniques of biological research. I created various clone libraries of Synechococcus DNA by isolating specific sequences extracted, amplified, and cloned through molecular methods. Through this, I was able to observe the relative preference of one “clade” of Synechococcus at deeper levels of water, suggesting the existence of adapted metabolic differences between Synechococcus found on the surface and those at deeper levels of water. Because this type of depth research on Synechococcus is relatively uncommon, I found these striking results very exciting.
Overall, my experience as a CCE LTER REU has opened my eyes to the exciting possibilities of biological research, as well as providing me with a greater understanding of difficult concepts from a hands-on perspective. I am looking forward to applying these skills to a future career in research science.
Kyra Rashid

This summer I had the honor of working in Dr. Michael Landry's lab where I investigated how climate change influences microplankton community distribution and dynamics. Under the guidance of third year Ph.D. student Ali Freibott, I spent the first two months of my internship processing samples that were collected along the frontal system from the 2012 CCE Process Cruise. Using light microscopy I calculated the biomass of the ciliate community and dinoflagellate community and then illustrated the distribution of these communities along the location of the frontal system. The data I procured will contribute to the overall study of microplankton community distribution and composition that has been compiled over the years of CCE LTER studies.
For the last month of my internship I had the amazing opportunity to live aboard the R/V Melville where I took part in multiple types of oceanographic sampling on the CCE P1408 cruise. I worked with a team of scientists to collect seawater samples whose analysis will help us understand microplankton community composition, distribution, and dynamics under El Nino-like conditions. In particular I became familiar with the design, execution, and analysis of dilution experiments which are a means of calculating community growth and grazing rates. We designed our experiments to explore the effects of size fractionation and nutrient limitation on phytoplankton communities. Additionally we made our first attempt to use metagenomics to calculate species specific growth and grazing rates.
My journey on the R/V Melville has been an invaluable living and learning experience. I am so grateful for the opportunity to work amongst such accomplished scientists. It was refreshing to learn about oceanography in such an interactive way. My incredible experience at Scripps this summer has inspired me to continue pursuing a career in the field of biology.
Jasmine Tan

This summer, I worked in Farooq Azam’s lab under the mentorship of Post-doc Byron Pedler Sherwood. My work as part of the CCE LTER REU program involved measuring protein synthesis rates of bacteria and converting synthesis rates into bacterial carbon productivity via leucine incorporation and centrifugation. I also participated in the CCE LTER P1408 cruise by taking seawater samples at different stations and depths off the coast of Point Conception. Before embarking on the process cruise, I created a project to test the best forms of storage on the cruise. I took seawater samples from the SIO pier, processing all samples within the same day, and studied the effects of storing samples across different periods of time and at different temperatures. With the results I collected for the duration of the month prior to the cruise, I learned that the longer a sample is stored and the higher the temperature it is stored at, the higher the variability. I used this information to my advantage while on the cruise.
My experience in this program has been eye-opening in more than just areas of science. I had the opportunity to sail into the middle of the ocean for a month, work on a research vessel, and meet people I thought I would have never met before. Working with SIO gave me a plethora of knowledge from other labs as well as my own. This internship has peaked my curiosity in not only biology, but other sciences and worldly topics as well. Most importantly, I learned the importance of bacteria, something I never thought I would be studying, in our lives, as it represents a major pathway for organic matter in marine ecosystems. In short, sometimes it’s the little things that matter most!
2013
Ana-Patricia Lopez

I am working with Art Miller and his research team this summer 2013; my work as part of the CCE LTER REU involves the quantification of spatial structures of phytoplankton and their relationship to ocean currents, waves and ocean fronts through data manipulation of large data sets generated with ROMS (Regional Ocean Modeling System) to learn whether the simulations resemble observations and to diagnose the physical controls of the system. The ecosystem model is unique in that it contains a large number of phytoplankton “species” which are given arbitrary characteristics that have trade-offs in their growth rates and physiological parameters. I am gaining very valuable skills as I am learning how to use MATLAB to read the netcdf files, in addition to learning about physical oceanography, which I did not have much background in before I started the internship program.
I have had the opportunity to meet other students and have learned from their grad student journey at SIO. Learning about the physical attributes affecting the biological ecosystem in the California Current System has been an exceptional learning experience as part of my environmental engineering preparation. I am so happy to be part of the team and I look forward to possibly continuing my graduate education at this wonderful institution. In addition, my goals are to have a better understanding of the subject as a whole, and being able to come to meaningful observations that will be a significant contribution in the field.
Anna Cassady

While working in Brian Palenik’s lab during the summer, under the mentorship of Dr. Palenik and Post-doc Javier Paz Yepes, I expanded upon work I had done previously there with different strains of Synechococcus cyanobacteria. During the spring I had worked studying the effects of secondary metabolites in the supernatant from each strain on the development of other strains. Throughout summer, I studied the effect these secondary metabolites had on sea life communities off the SIO pier. This was accomplished by setting up half of the experiment to include seawater with the supernatants of four strains studied. In addition, I set up the other half with seawater and the cells of the same four strains. The experiment is monitored by utilizing both microscopy and flow cytometry. With the first half of the experiment, I will be able to study whether or not the abundance of different genera are effected, and with the second part I can see what the effect, if any, had on the grazer community.
Alicia Highland

As part of my REU experience, I have the opportunity to work under the guidance of Dr. Mark Ohman and second year Ph.D. student, Jennifer Brandon. My research focuses on mesozooplankton grazing rates across oceanic fronts in the California Current Ecosystem, with major emphasis on pelagic copepods. Copepods contribute vitally to both oceanic food webs and the global carbon cycle, therefore understanding how they effect and are impacted by changing oceanic conditions is becoming increasingly important.
The California Current Ecosystem experiences natural variability at seasonal, annual, and decadal scales. However, long term research conducted by Dr. Ohman and his colleagues indicates that the CCE is experiencing increasing and continual warming as a result of global climate change. Their research also indicates that as ocean winds and other conditions change, the occurrence of submesoscale ocean fronts is increasing. Ocean fronts can be “hot spots” for primary production and provide extraordinary feeding opportunities for species at various trophic levels. The aim of the research that I am conducting this summer is to determine whether zooplankton grazing upon phytoplankton changes along these frontal boundaries. In order to do so, I am utilizing the gut fluorescence method.
The gut fluorescence method estimates the grazing rate of zooplankton by quantifying the amount of chlorophyll-a and phaeopigment in the zooplankton pre-egestion. This is accomplished initially by sorting the collected zooplankton from residual phytoplankton and placing them in an acetone solution. The chlorophyll pigments are then released from the zooplankton tissue using sonication and centrifugation. Finally, the resulting supernatant is analyzed with a fluorometer.
Being that I am an Ohioan, born and raised, having the opportunity to be part of the CCE LTER program and conduct meaningful oceanographic research is truly incredible. Through this experience, I am learning valuable research methods that I hope to employ during my graduate studies. I am also learning a great deal about the graduate school process, the ins and outs of maintaining a lab, and the importance of collaborative effort. Most importantly however, I now have a greater understanding of zooplankton ecology and appreciate the very important role zooplankton play on both local and global scales.
Emma Nuss

Throughout my senior year attending UC San Diego (2012/13), I was given the opportunity to participate in the CCE-LTER REU program. Working with Dr. Art Miller, I spent the year looking at the output of a Darwin model. My goal was to investigate the relationship between biological activity in the ocean and physical processes, in particular eddy currents. The main question that I was trying to answer through my research was what happens to phytoplankton in an eddy? To answer this question I spent most of the year making various plots of the 20 different phytoplankton variables in the Darwin model and looked at their concentration in one eddy off of the Southern California coast. I looked into their distribution in relation to depth as well as through time. Towards the end of the year I began looking more specifically into the correlation between different phytoplankton variables and surface height.
By the end of the year I became quite familiar with Matlab plots and netcdf files. My experience working with Art Miller developed my skills as a scientist and his advising style encouraged me to ask relevant questions to keep pursuing my research questions. Overall my time in the CCE-LTER REU program has been a great experience; it has given me a sense of what research in Oceanography is. The skills I have learned over this past year have helped me prepare myself for my future studies in Oceanography in graduate school next year!
Jenny Couture

For my internship in the CCE-LTER REU program, I had the opportunity to work with Dr. Tony Koslow on a project that investigates our ability to assess the state of the ocean. More specifically, I carried out a meta-analysis on Pacific Ocean observation programs to determine where and how the physics, chemistry, and the biology of the ocean are measured. From this analysis and through mapping these parameters, Dr. Koslow and I determined that even though the observation programs are measuring the physics and the chemistry of the ocean adequately, biological monitoring is severely lacking or inconsistent. This issue is of particular importance today because scientists are unable to accurately evaluate the state of the marine ecosystem as a whole.
Overall, my experience with the CCE-LTER program has been invaluable. This wonderful opportunity has enabled me to learn about the field of oceanography and the scientific research each Pacific Ocean observation program is currently conducting. In addition to learning about these various monitoring programs, I also had the chance to attend research seminars where I became familiar with the diversity of research at Scripps. I am excited to apply what I have learned this summer in my future career in the field of biological sciences.
2012
Aarani Arulmoli

For my internship in the CCE-LTER REU program, I had the opportunity to work with Dr. Kathy Barbeau and the graduate students in her lab to study the role bacteria play in the biogeochemical cycling of iron. My project involved metagenomic analysis of the microbial community in environmental samples and in iron addition grow-out incubations. In order to construct a clone library for microbial community analysis, a target gene had to be selected as the phylogenetic marker. Since the 16S rRNA gene contains both highly conserved and variable regions, it was chosen as the appropriate marker. The target gene was then amplified from environmental DNA extracts using PCR. I learned how to purify these amplicons and insert them into a plasmid vector. I became very familiar with the E.coli cloning procedure to grow out the competent cells with the recombinant vector and how to purify these plasmids with the gene insert to prepare them for analysis. I learned how to use sequence databases and alignment tools along with how to make phylogenetic trees to evaluate bacterial community composition.
This opportunity to participate in the REU program has been an extremely valuable learning experience. This summer I have not only gained familiarity with using techniques in microbial and molecular biology, but I have also learned the importance of perseverance and patience when conducting research as it can be a slow and challenging process. The program also gave me the unique opportunity to hear the perspectives and advice of several graduate students based on their personal experiences. The skills and knowledge I have gained throughout this summer will definitely carry over into my future pursuits in graduate school and beyond. I am especially grateful for the guidance I have received from all the members of the Barbeau lab and am excited to continue and expand my research during this coming academic year as part of my ESYS senior internship.
Carole Wang

This summer, I interned in the Palenik lab, which focuses on studying a marine cyanobacterium called Synechococcus, and also volunteered with the Landry Lab on the CCE process cruise (P1208). On land, I had the great opportunity to learn about a variety of topics with my fellow undergraduates under the supervision of Dr. Brian Palenik, including sustainable biofuels and seasonal green blooms. My specific project focused on Synechoccocus clumping. Synechoccocus are traditionally thought to be unicellular phytoplankton, and the recent discovery that they can (and consistently do) form micro-colonies in the field is opening the doors for exciting new research. I introduced external stressors in the form of sugars and other chemicals to Synechoccocus cultures to test whether these chemicals can induce or disrupt clumping. The samples were analyzed via epifluorescence microscopy and a grid counting chamber, while cell growth was monitored through fluorometer readings. The purpose was to develop an idea of what sort of structures exist on the cell membrane surface that may be involved in this clumping behavior.
During the CCE-LTER research cruise, I assisted in collecting water samples from the CTD (Conductivity, Temperature, and Depth recorder) Rosette and from drifters. The CTD Rosette collects water from several depths, while the drifters are used as a method to incubate phytoplankton and grazers within the ocean. These samples were filtered, while some were preserved in liquid nitrogen for future analyses on POC, chlorophyll, and DNA content. Other samples were analyzed for chlorophyll content on the ship using a fluorometer. I also collected some samples across a front to observe Synechoccocus clumping distribution. These samples were preserved with paraformaldehyde and stored in the -85˚ freezer and later made into slides to be observed via epifluorescence microscopy.
It was rather odd working through the night and eating breakfast for dinner, but that also became a part of this unique and memorable adventure. This summer was packed full of interesting experiences that, a year ago, I wouldn’t have thought to be available for undergraduate students. I am grateful for this amazing opportunity, as it has given me several fresh new perspectives and has also helped me develop a better understanding of what I’ll be doing in the future.
David Muller

Over summer 2012, the CCE-LTER REU program gave me the opportunity to work in Dr. Todd Martz' lab here at Scripps. One goal of the Martz lab is to investigate the effects of ocean acidification. Accordingly, the lab specializes in developing autonomous chemical (especially pH) sensors for the ocean - the SeapHOx instrument package being one of their hallmarks. The SeapHOx returns especially valuable information about ocean pH because of its autonomous design, as it sits on the sea floor and takes measurements every half hour over the course of several months (many more measurements than you could take from a ship).
My project for the summer involved updating the outdated computer and overhauling the software that runs the SeapHOx. I spent most of my time writing software in C to interface with the dissolved oxygen sensor, salinity sensor, 24 bit analog to digital converter, and pH sensor on board the SeapHOx. Putting all these instruments together in one place and running them off of one computer proved to be no small task! I learned a lot about electrical engineering/digital circuit design and honed my software skills putting all the code together.
I think the experience of working at Scripps was just as valuable as the work itself. It was really meaningful for me to get a glimpse into the life of graduate students in a lab setting (especially one as picturesque as Scripps). It has definitely piqued my interest in pursuing a higher degree in just about any of the sciences. Overall, the REU experience has been overwhelmingly positive - I'm really grateful that I had the chance to work at Scripps for the summer.
Dustin Chen

Hello everyone! My name is Dustin Chen, and this past summer I did a CCE-LTER REU internship under the guidance of Rebecca Asch (Ph.D. student) in the lab of Dr. Dave Checkley. For my project, I chose to test the validity of a hypothesis called the “school trap.” This hypothesis states that, when a schooling fish species reaches population levels so low that it can no longer form schools by itself, it will start to school with another species of schooling fish. This can be detrimental to the first species because the other species may not occupy the same habitat or may compete against it for food. My study focused on the following three fish species: Pacific sardine (Sardinops sagax), chub mackerel (Scomber japonicus), and jack mackerel (Trachurus symmetricus).
I used MATLAB to analyze a dataset containing egg and larval concentration data spanning from 1951 to 2010. I investigated how historical habitat overlap between sardine and either mackerel species correlated with three proxies for sardine population levels: egg and larval concentrations, percent station presence, and spawning stock biomass. Correlation analyses indicated that the school trap may be occurring between sardine and chub mackerel, as the relationship between increased habitat overlap and decreased spawning stock biomass was highly significant.
I also began a diet comparison study of locally collected Pacific sardine and chub mackerel. This study is still in progress.
Overall, being a CCE-LTER REU intern was an invaluable experience for my growth as a scientific researcher. I was able to learn how to use MATLAB in scientific research and basic statistics, and how to perform a fish gut content study. Even more than that, I was able to see firsthand what I am capable of accomplishing within a limited timeframe. This deepened my understanding of the incredible amount of time and effort put into carrying out a research project.
Jonathan Cheng

For my summer REU at SIO, I worked in the lab of Dr. Farooq Azam, under the guidance of Ty Samo, a PhD student. I studied marine bacteria and the role they play in the biogeochemistry of the ocean. My main focus of the summer was to measure the rate of bacterial carbon production, using a radioisotope incorporation technique. Like most studies involving bacteria, there was also lots of microscopy involved with my studies. I analyzed samples from the previous CCE process cruise (2011) before I left on the R/V Melville for a 30-day (2012) process cruise. While on the ship, I gathered and filtered samples for bacterial community growth analysis, and froze some for making slides back on land. In addition, I helped with other projects.
The cruise was an awesome experience. I consider myself extremely lucky to have participated in such a project. I got to see and do so much of the things I have only heard about in class. Also, just being around so many graduate students and PI’s was an ideal learning experience, and something that I really cherished. My whole summer, in the lab and on the ship, was great. Of course I learned a great deal about microbiology, and expanded my love of the ocean, but I learned about how science gets done in the real world.
2011
Ellen Umeda

For my CCE-LTER internship, I worked with both Dr. Mark Ohman (Lead PI) and Linsey Sala, Assistant Museum Scientist at SIO. My research experience began with a month long cruise on the R/V Melville studying fronts off the coast of Point Conception. I helped with the deployment and recovery of the bongo and MOCNESS tows. When the bongo net came back on deck, I quickly anesthetized the zooplankton sample with club soda, size fractioned the sample, and placed it in liquid nitrogen. These samples are for gut fluorescence and biomass analysis, which help determine the grazing rates of zooplankton. The MOCNESS consisted of ten nets, making it look like what I called the "Mocness Monster." Each net could be triggered to open at a specific depth. Washing down each of the nets and preserving all ten samples took a lot of time, but we caught a lot of interesting animals including hatchet fish, pyrosomes, jellyfish, dragonfish, and a solitary salp. For me, the highlight of the cruise was seeing a large pod of Pacific White-sided dolphins. After completing my first research cruise, I learned about life out at sea and had the opportunity to meet other professors and students interested in marine research.
Back on land, I compared vertical distribution and life history stages of pyrosomes. I sorted through almost 100 jars from the MOCNESS tows and pulled out all of the pyrosomes. I then measured the pyrosomes, which ranged from 2mm to 288mm, and calculated the density of pyrosomes using the volume of water filtered by each net. The pyrosomes appeared to show a vertical migration toward the surface at night. In addition, the data from the day tows showed that larger pyrosomes occupy a greater range of depths including deeper ones.
This REU internship gave me the opportunity to apply what I have learned in classes to oceanographic research. I never realized how much more goes into research than simply reading textbooks. From this experience, I learned how to collect samples on a research cruise and analyze the samples back on the land. This experience has encouraged me to pursue a higher degree in marine biology.
Evelyn Hunter

For my REU internship, I worked with Dr. Ralf Goericke (Co-PI) to manage data from the CCE LTER cruises. My primary project was to create a database which included data from the ALF (Advanced Laser Fluorometry) device and underway data. I spent a lot of time writing and running functions in MATLAB to consolidate and organize numerous smaller text files into one complete file, then importing those text files into a Microsoft Access database. I then created queries in the database so the content could be searched with different criteria- location, time, and other measurements. When I wasn’t in front of a computer, I was calibrating fluorometers at yacht clubs, loading the research cruise ship and attending seminars put on by the SURF program.
My time at SIO was a great chance to learn new material as well as become more familiar with applications I already knew. I am so grateful for the learning opportunity in this institute; seeing all of the different divisions of scientific research in action and their possible applications was really exciting and interesting. As I continue my education, I will undoubtedly use the knowledge and research experience I gained through this program.
Kayla Blincow

This summer I interned in Dr. Tony Koslow’s lab. I have been volunteering in Dr. Koslow’s lab for a few years now, but this CCE-LTER REU gave me the opportunity to expand on the skills and lab experience I have been developing as a volunteer. I was able to analyze the data I have been collecting and start a project of my own. I was also given the opportunity to be the Oozeki technician on the summer 2011 CalCOFI Cruise. I gained invaluable experience at sea, working with other researchers, learning different sampling techniques, and getting a taste of life on a research vessel.
The project that I worked on this summer, and plan on continuing to develop in my final year of undergraduate studies, involves analyzing data collected from the Oozeki net on the CalCOFI cruises to map euphausiid communities in the California Current. In the lab, I identify euphausiids to the species level and count the number of each species from the cruise samples. With this information it is possible to estimate the abundance of each type of species, and map their community structure based on where the sample was collected. Ultimately I would like to identify environmental and/or ecological factors that might be contributors to the abundance patterns that appear. Frontal, temporal, and spatial changes are all likely to have an impact on euphausiid assemblages in the California Current.
I intend to get my PhD in marine science, and this internship introduced me to experiences and people that will help me achieve that goal. I am extremely grateful to Dr. Tony Koslow and Post-doc Dr. Ana Lara Lopez for mentoring me this summer and over the past few years. This summer’s REU taught me a lot about marine research and reaffirmed that it is a career path I intend to pursue.
Lisa Della Ripa

My CCE-LTER research project involved comparing and assessing the trophic positions of zooplankton (0.2-2 mm) in three different biogeochemical environments: California Current Ecosystem (CCE), Costa Rica Dome (CRD) and Hawaii (ALOHA). This research is important because fisheries make models to better manage stocks, which rely on the trophic positions of fish in the food web. These fish feed on zooplankton and phytoplankton, which make up the base of the food web. We were investigating if there was a 3.4‰ (permille) increase of δ15N between the 0.2-2 mm size fractions as models suggest. We also wanted to know if there was a difference in δ15N in the source nitrogen between the different regions and if the absolute trophic positions are different between regions. This internship also involved participating in a month-long research cruise off the coast of Point Conception in California on the R/V Melville to collect the CCE samples. I analyzed all the samples in Dr. Brian Popp’s lab at the University of Hawaii at Manoa using both Bulk Nitrogen Analysis and Compound Specific Isotope Analysis (CSIA) methods. While I was in Hawaii, I also participated in a six-day research cruise at station ALOHA. I learned a lot about oceanography on these cruises and have truly had an exciting and adventurous summer!
This opportunity couldn’t have come at a better time, since I recently switched my major from biology to chemistry, and this internship allowed me to conduct research at the interface of chemistry and biology in a marine setting. I have learned so much this summer, gained valuable field experience, lab techniques and data analysis methods. I thank my mentors - Dr. Mike Landry, Graduate student Moira Decima, and Dr. Brian Popp - for making this a very productive and exciting summer. This research opportunity has encouraged me to further explore the world of marine science!
Melinda Lopez

For the 2011 summer CCE-LTER REU program, I was given the opportunity to work in Dr. Todd Martz’ lab, which constructs an autonomous instrument package called the "SeapHOx." My background in physics and computer science helped me to create additional software functionalities for the program running on the SeapHOx’s embedded microcontroller. I was able to program, calculate, and display the pH by using certain measured or user input pH calibration sensor coefficients, such as temperature and salinity. I also created a Lab View program for interfacing the microcontroller with a PC.
Before the start of the summer my knowledge of chemistry was limited to the basic 101 course and my skills in programming were never put in practical use beyond class room exercises. I spent a great deal of the summer troubleshooting computer programs and trying to get the sensors to do exactly what I wanted. Even though this task was very difficult at times, it helped me hone in and expand my computer programming skills. Spending the summer in the Martz lab also allowed me to see how interdisciplinary the sciences could be, and to see what kind of research other scientists outside my field of physics conducted.
Wendy Plante

For my summer REU internship, I worked with the Barbeau lab (led by Dr. Kathy Barbeau, Co-PI) studying the role of Iron in the marine ecosystem. This included the opportunity to participate in a one-month oceanographic research cruise on the R/V Melville. I got to meet many different people interested in many different fields of science. It was great to see all of the different research projects coming together to study this water system. The all-night sampling sessions and early morning casts were definitely something I had to get used to, but ended up nostalgic about when the cruise came to an end. During the cruise I assisted in casting the trace metal clean rosette, cleanly filtering the samples collected, and running experiments examining how Iron affects the prevalence of primary producers.
The rest of my time at SIO was spent examining the samples collected on the cruise. Specifically, I ran Iron (II) lifetime experiments, comparing the oxidation rate of Iron (II) in each water sample collected in the CCE study area. The water that I used was taken from off the coast of Point Conception. There were two surface samples, each taken from either side of the front that we were studying, and one taken from 250m at the further off-shore station. Decay experiments were run by analyzing the concentration of Iron (II) over time using an FeLume. The FeLume measures the amount of light produced by the reaction of Luminol with any Iron (II) in the seawater sample. The graph of the natural log of the chemiluminescence signal during the decay was then used to obtain the oxidation constant and find the half life.
This summer provided me with a unique perspective into the lives of oceanographic researchers with experience in field work as well as lab work. I am truly grateful for all of those who assisted in this learning experience – whether it was through mentoring or by merely taking the time to discuss their own research experiences.
2010
Kayla Busby

For my internship in the CCE-LTER REU program, I spent the summer in Brian Palenik's lab, researching the seasonal and geographical diversity of Synechococcus sp. CC9311. Synechococcus is an abundant marine cyanobacterium that plays a significant role in global carbon fixation, and has also been shown to be more sensitive to high levels of copper than other marine phytoplankton.
During the course of this summer, I examined the environmental prevalence of an operon that is upregulated in Synechococcus sp. CC9311 in response to copper shock. It is also suspected that this operon may have been horizontally transferred from estuarine organisms. Using quantitative PCR (qPCR), I analyzed environmental DNA samples and quantified the amount of DNA corresponding to this operon, in addition to quantifying the amount of DNA corresponding to all Synechococcus sp. CC9311 organisms. With this technique, I was able to determine the relative abundance of this operon, both seasonally and geographically. I did this through the analysis of environmental DNA samples collected throughout the course of a year at the SIO pier, and environmental DNA samples collected at estuarine areas and other coastal sites.
Overall, this summer has been an amazing experience, and I have learned a lot - in terms of lab techniques and research skills, and also of my own future goals and aspirations. It has been a pleasure to work with everyone in the Palenik/Brahamsha labs, and I am especially grateful for the mentorship and thoughtful guidance of Post-doc Rhona Stuart and Dr. Brian Palenik.
Ross Castillo

The CCE-LTER REU summer program allowed me to work with Prof. Kathy Barbeau and her graduate students studying trace metal biogeochemistry. To better understand aspects of iron cycling in marine systems, I performed experiments on the SIO pier, PCR with degenerative primers, bioinformatics analyses of discrete organisms and metagenomic datasets, and determined antibiotic resistance for a model organism in preparation for gene knockout experiments.
In the wet lab, I learned to use instruments like the voltammeter and the flow injection analyzer to study iron chemistry in pier samples. To measure chlorophyll during our pier incubation experiments, I used a fluorometer. I learned how make liquid media and plates and grow bacteria on them. Studying the best antibiotics to use in knockout studies, I learned how to make these media with different concentrations of antibiotics and measure the growth of the bacteria using a spectrophotometer.
I combined wet lab experiments with bioinformatics when developing degenerate primers and identifying proteins of interest to study acquisition of heme as an iron source by marine bacteria. Degenerate primers are used to amplify a broad range of DNA sequences. To identify conserved sequences to develop these primers I used alignment tools. I then learned how to perform different types of PCR, such as gradient and touchdown, and run gels. To develop Hidden Markov Models, I learned to use statistical modeling tools to help perform more accurate searches against sequence databases and metagenomic datasets.
This educational opportunity has given me hands-on practice with important techniques that I will be using in my future research. I am grateful for the guidance that I have received during this program and the insights that I have gained as a result. However, what made this experience truly special were the friendships that I developed while working on the beautiful SIO campus over the summer.
Chelsea Didinger

This summer I researched zooplankton in Dr. Dave Checkley's lab. I had the opportunity to participate in an 18-day research cruise to Monterey Bay aboard the R/V New Horizon. On the cruise, I worked the noon to midnight shift and was able to see an assortment of marine life, including Humboldt squid and humpback whales that came within a few feet of the ship. I assisted in the deployment and recovery of instruments, such as the CTD, MOCNESS and Bongo-LOPC and helped preserve the samples collected. With the help of my mentor, I also designed and ran an experiment to measure the excretion rate of euphausiids.
Back in the lab at Sverdrup Hall, we processed data from the cruise. The programming software Matlab was used to analyze the depths that the MOCNESS nets opened by correcting for sigma-t values read by the CTD and the MOCNESS. I joined the Scripps collector in gathering copepod samples and then measured their excretion rate. To calculate this value, a fluorometric method is used in which the main ingredient in the working reagent reacts with the nitrogen in ammonium and fluoresces. The raw unit readings from the fluorometer allow for the calculation of very low ammonium concentrations.
This internship was an unparalleled opportunity. I was fortunate to meet new associates and friends, to learn how to design and refine my own experiments, and to participate in oceanographic research firsthand. This experience reaffirmed that studying at Scripps Institution of Oceanography is the path I intend to follow.
Danielle Lara

For my REU internship, I worked in Dr. Lihini Aluwihare's lab at SIO. This past summer, I helped determine the level of carbohydrates in seawater samples taken from the CCE LTER 2008 cruise. The concentration of carbohydrates was determined using a sensitive colorimetric assay. Referred to as the TPTZ method, it utilizes specific reagents to react oxidized sugars to promote the formation of a colored complex. The absorbance of each sample can be taken using a spectrophotometer, giving some indication of the carbohydrate level of each station in the ocean. Because the samples were taken at various depths of the water column, a trend of carbohydrate concentration was seen, depending on sample station.
This experience has taught me a lot about the ocean but most of all, about myself. It taught me that colorimetric assays are, in fact, a trying and tedious method to obtain results. Every assay was run with as much precision as possible but due to the many outside factors that can affect the results, it was somewhat difficult to obtain a baseline for even the reference deep sea water sample. I learned that regardless of tribulations, it is important to keep going and push forward. After all, it is research! I greatly enjoyed my time and summer with the members of the Aluwihare lab, due to all of them and their welcoming and helpful nature. This experience will stand as one of the most influential and rewarding events of my academic career.
Emilie Schnarr

As an REU student, under the guidance of Graduate student Katharine M. Hanson, I analyzed the number, diversity, type, and size of zooplankton consumed by the yellowtail damselfish, Dascyllus flavicaudus, sampled from various reef habitats surrounding the island of Moorea (another LTER site - MCR). Through microscopic investigation of 20 stomachs, each containing approximately 2000 individuals, oceanic and reef-associated zooplankton were compared. It was observed that the yellowtail damselfish consumes a wide range of prey items, including several copepod species, fish eggs, appendicularians, and meroplankton.
Prey item body-size measurements were used to determine the quantity of carbon consumed by the yellowtail damselfish and to distinguish between oceanic and reef-associated zooplankton. The ZooScan with ZooProcess (M. Ohman's lab) was used to determine prey item size. Each prey item was scanned to obtain automated ZooProcess body-size measurements. ImageJ software was used to manually measure body size. A relationship between automated measurements and manual measurements was constructed for each prey category and thus ZooProcess calibration will allow for future prey item body-size measurements.
My REU experience this summer has been very rewarding. I have learned many lab skills and have met many interesting people. The research should provide insight into complex food web dynamics across reef ecosystems on the island of Moorea. This year, as a senior thesis project, I expect to complement this research by conducting a feeding electivity study for the yellowtail damselfish.
2009
Nadija Alainentalo-Anderson

In the CCE-LTER REU summer program, I worked in Dr. Kathy Barbeau's lab to investigate the role of iron and silica as limiting nutrients for phytoplankton growth in the California Current Ecosystem. My research experience included analyzing samples from past cruises in the CCE LTER study site as well as conducting my own "fieldwork" via nutrient-amended incubation studies on the Scripps pier. The incubation studies performed on the pier allowed me to learn trace metal clean sampling techniques to prevent contamination and to participate in cruise-like sampling. During incubation experiments I took daily microscopy, nutrient and chlorophyll samples. Samples were also taken for pigment analysis and iron speciation analysis. I used a fluorometer to measure chlorophyll.
The main laboratory analysis I conducted during my internship involved using high performance liquid chromatography (HPLC) to identify and quantify phytoplankton pigments. These pigments are indicative of certain phytoplankton species and were used to characterize phytoplankton community structure in seawater samples. These seawater samples were taken from various incubation experiments conducted on cruises in the CCE LTER study region by members of the Barbeau lab, as well as samples collected during my summer 2009 pier incubations. In support of the HPLC data, preserved seawater samples were examined on an inverted microscope to confirm community composition. I am continuing this research during the school year on a part-time basis. I have learned a number of new laboratory techniques and feel that this experience has allowed me to make an informed decision about my ambitions to pursue oceanographic research in graduate school.
Emily Botts

For my REU internship, I worked in Brian Palenik's lab. The primary focus of this lab is on the marine cyanobacteria, Synechococcus. Synechococcus is a diverse genus of phytoplankton that is an important primary producer in the oceans and contributor to the planet's carbon cycle. My project was to compare the diversity of coastal versus offshore populations of the cyanobacteria from a CalCOFI cruise using the high-resolution melting technique.
In high-resolution melting, DNA is extracted from filters and amplified using PCR. A fluorescent dye is used during the PCR, which would only fluoresce when the DNA is in a double helix. DNA separates or melts into two strands when it is heated and the melting temperature depends on the GC base pair content and sequence. The fluorescence is recorded as the DNA is slowly heated and the data points produce a curve from which a unique melting temperature for each present strain can be determined.
The opportunity to participate in this program has been a very valuable experience for me, as someone who intends to go on to professional school for biological research. Being able to work in an actual lab on a daily basis allows for newly acquired lab techniques to be practiced and perfected in a way that is not reproducible in the classroom.
James Hayes

I worked with materials from twenty-six long-term ecological research sites for my summer internship. This effort yielded a corpus of materials about information management that will support research on the development of site-based infrastructure. My work involved documenting information systems development, data handling, as well as changes associated with technology and with the roles of information management. A survey of the sites led to highlighting commonalities and differences among the sites and at individual sites over time. I worked in the Ocean Informatics Design Studio alongside informatics specialists, information architects and programmer analysts.
As a student of the history of science it was enriching to experience a site contributing to the development of information infrastructure and information management. No amount of time with texts would have afforded the same chances to witness science as a living enterprise. In addition to the opportunity to observe informatics and research professionals in their milieu, I’ve learned technical skills and become familiar with hardware and software that had heretofore rarely been a part of my formal education. SIO may only be a couple shuttle stops from main campus, but this program prompted me to consider the history of science from a more privileged and broader perspective.
Patricia Lee

For my REU internship, I worked in the Aluwihare lab and helped the quantification of the total organic carbon (TOC) content in the samples that were taken from the CCE LTER 2008 cruise. The amount of TOC is determined by running the samples through a TOC machine. In the TOC machine, a high-temperature catalytic oxidation (HTCO) method of measurement was conducted. Through this method, the existing inorganic carbon is removed. Then, the carbon that’s left in the sample is combusted to CO2, which is quantified in a nondispersive infrared sensor.
In this experience, I was able to learn a lot of trouble shooting skills. It was difficult to get precise and accurate data at the beginning of my internship and understanding the machine was a key to making modifications to increase the quality of the data that was obtained through the machine. After many weeks of alterations to the machine, the processed samples helped me conclude that there are high levels of TOC towards the surface of the ocean and lower levels as the depth increases.
Junna Murakawa

As an REU intern in the Ohman lab at SIO, I studied the grazing rates of meso-zooplankton. Mesozooplankton are zooplankton within the size range of around .2 mm to 5mm that consume other plankton. The amount of chlorophyll a pigments from phytoplankton remaining in the guts of mesozooplankton is used to determine the grazing rate. This can be analyzed by looking at the level of fluorescence of the extracted gut pigments. The samples were taken in the California Current Ecosystem during an October 2008 cruise. Samples were taken at various locations and times. By analyzing the gut pigment contents we can look further at the differences in grazing rates in relation to proximity to shore, different size fractions, diel periodicity, and other factors.
From this summer internship I have learned a lot about an area of science that I wish to continue studying in the future. I was able to gain experience with lab techniques, data entry and analysis. I worked with Dr. Mark Ohman and with his guidance learned a great deal about many aspects of science. I also learned about plankton, their grazing capability, and roles these microscopic organisms have in the marine ecosystem. Although I am unsure as to what specific area of marine science I want to pursue, this internship has given me an opportunity to discover and experience in detail one specific area of marine science. This internship has been an irreplaceable experience that I will definitely carry on into my future.
Robert Peterson

The REU experience this summer afforded me an opportunity to engage with informatics, work that lies in the overlap of information science, social science and oceanography. Hands-on experience in a contemporary information environment is key to developing as an environmental sciences scholar today and to participating in "big science". As part of the LTER network, the Baker lab's approach involves collaboration with local, community and network entities to further the development of knowledge. The philosophy of continuing design and active engagement with that process are key to how the Ocean Informatics team addresses the long-term issues involved in knowledge-making. Continuing design is a way of carrying out information management in support of scientific research when both are in active development.
My particular REU projects involved implementing a geographic dictionary or gazetteer and initiating development of a history module that incorporates a timeline of events designed to enable the capture of project narratives. The day-to-day software development as well as the longer-term project planning and collaboration efforts grounded the abstract practices only touched upon in the classroom. The interdisciplinary nature of the lab gave me a chance to meet and engage with scientists in other fields. Working with the staff in the lab provided valuable perspectives that would otherwise be unknown and unavailable to me. The lab's engagement in the academic areas into which I rarely ventured such as oceanography and social science help to create in me a broader and uncensored understanding of how the work with scientific data really gets done.
Tim Stillinger

Hey! My name's Timbo and for my research this past summer I worked on correlating in situ acoustical backscatter data of zooplankton samples collected with an Acoustic Doppler Profiler (ADP) to zooplankton biomass estimates. Most of my time in the lab was spent digitizing zooplankton samples with Zooscan. Zooscan involves a waterproof scanner that captures high resolution images of hundreds of zooplankton at once. Zooscan then separates each zooplankton scanned into individual images and computes various measurements for each individual. I classified each individual zooplankton sample by species (over 150,000!) and from the measurements and species identification computed the total biomass of the sample. I then compared this biomass data to the ADP backscatter data to determine any correlation between the two. This was all preliminary research for SIO graduate student Jessie Powell's thesis.
I also went out into the field for 21 days onboard R/V New Horizon on the SEAPLEX cruise to the North Pacific Gyre. On the cruise I helped with the scientific deployments and worked the night shift from 8pm to 8am. It was my second time out to sea on a research vessel and I had a blast! On this cruise I also helped Jesse Powell deploy Bongo Nets with an ADP installed, which we used to collect more samples to be analyzed in a similar fashion to the samples I analyzed earlier in the summer.
Overall, I had a wonderful time working in the Ohman lab this summer and hope to be a graduate student at SIO in the very near future.
Elliot Weiss

During the course of my REU internship, I performed research on phycobiliproteins, which are water-soluble photosynthetic pigments present in cyanobacteria, cryptophytes, and planktonic rhodophytes. Within these organisms, the phycobiliproteins are used as accessory pigments to absorb light at wavelengths which are not utilized by chlorophyll. When exposed to light, phycobiliproteins have distinct excitation and emission maxima. The specific wavelength of this maximum is altered depending on which type of organism the phycobiliprotein is contained within. Using this knowledge, one can determine the phycobiliprotein-containing community structure of a given water sample using fluorescence.
Over the summer of 2009, I collected water samples from the Scripps pier and stored them in liquid nitrogen. By implementing and modifying a phycobiliprotein extraction technique established by Stewart and Farmer, I was able to extract the phycobiliproteins from these samples and use fluorescence to determine their presence. By observing the relative fluorescence of the samples over the course of the summer, I was able to observe changes in the coastal community structure. The main phycobiliprotein-containing organisms were cyanobacteria, which are an extremely important part of the marine phytoplankton community as they are important primary producers, especially in open waters. The method practiced was then used to validate the Advanced Laser Fluorometer, a new instrument designed for efficient shipboard analysis of the presence of phycobiliproteins and chlorophyll.
Overall, this was a tremendous learning experience. With the guidance of Dr. Greg Mitchell, my eyes were opened to the complexity of phytoplankton communities, and I learned a great deal about the nature of pigments. I also learned many lab techniques and data analysis methods which will be of great help to me during further education.
Sean Wiley

The Ocean Informatics Lab combines Oceanography, Information Sciences, and Social Sciences. Dealing with data collected by scientists and creating an accessible way for data to be accessed and shared among the community, the local information system called Datazoo creates an opportunity for learning in a research environment. During my REU internship I learned a number of programming languages such as MySQL, XSLT, XQuery, XPath, and PHP. Much of the knowledge I gained about these languages came from studying example code and researching online, which is a different approach than that normally used in a classroom. I also learned techniques and best practices through implementing designs, working collaboratively as part of a design team, and participating in discussions of published papers at weekly Informatics reading group meetings. I was able to take part in designing and implementing programs that deal with information and data associated with the LTER Network. Besides the backend software, my REU experience has also enriched my interaction skills because of the consistent feedback and teamwork provided by a diverse group of peers.
I entered the internship as a second year Computer Science undergrad. I had knowledge of languages such as Java, C++, C, and been exposed to others such as PHP, Pearl, and JavaScript. My presumptions were that I would be putting the skills I already had to use in the lab, so the breadth of information that I learned was unexpected. Working with the Ocean Informatics Team in Karen Baker's lab has allowed me to gain invaluable hands-on experience with not only the technical aspects but the community and communication aspects as well. During my research experience I gained the technical skills to program backend databases/software as well as interact with the users of these databases/software. Such applied knowledge will undoubtedly assist me in my continuing studies at UCSD and my future career.
2008
Todd Langland

For my REU internship, I performed research on zooplankton using the Zooscan, a waterproof scanner that makes digital images of plankton samples. More specifically, I studied the relationship between size measurements provided by these scans and the carbon and nitrogen content of the organisms. The purpose of the study was to provide a useful link between the new imaging and sorting technology of the Zooscan and more classic measures of biomass and organism makeup. 
Samples were collected during near shore tows just off the coast of La Jolla and brought back to the lab weekly. Lab analysis involved scanning and identifying organisms, followed by analysis of carbon and nitrogen values. The organisms in the scans were then hand measured using a computer and compared to both program measurements and their elemental composition. I then learned how to organize, graph, and analyze the data.
This experience has taught me a lot. I learned what it is like to work in a lab and about specific techniques related to what I was doing, data analysis methods, and a whole lot about what zooplankton look like and how they behave. Working with my mentors Alison Cawood (graduate student) and Dr. Mark Ohman (CCE lead PI) has given me a much greater understanding of this subject and provided me with a great opportunity for future education.
Mary McCormick

I have been researching how changing environmental factors alter food webs in the California current. I hope to determine whether organisms change their feeding habits, or if the makeup of their food is changing over time. I approached this by analyzing the nitrogen isotopes in tissues of three species of small animals that are common in the California current. Specifically, I selected samples of copepods, euphausiids, and sardine larvae that were collected between 1997 and 2000 off Point Conception as part of the ongoing California Cooperative Oceanic Fisheries Investigation (CalCOFI) cruises. The time series include the 1997-1998 El Nino, a period when conditions were warmer and there was generally less coastal upwelling to deliver nutrients to the sea surface, as well as the ensuing La Nina, during which ocean temperatures in the region tended to be cooler and increased upwelling delivered more nutrients to the surface.

My guides in this CCE REU project have been Dr. Mike Landry and PhD candidate Moira Decima at SIO. The analysis procedure I used, called amino-acid specific stable nitrogen isotope analysis, reveals the ratio of two Nitrogen isotopes (14N and 15N), which depend on an organism's eating habits and is therefore used as an indication of trophic level. This was conducted in the laboratory of Brian Popp, and also made possible by his and Elizabeth Gier's expertise) at the University of Hawaii.
Lisa Kamino

Dissolved organic nitrogen (DON) is the most abundant form of N (inorganic and organic N) accumulating in many areas of the surface ocean. In the CCE, DON dominates the total nitrogen reservoir during some summer and fall months. While much is known about the cycling of inorganic N, little is still known about the cycling of organic N in marine environments. For example, are its sources identifiable? Is it available to fuel biological production? Is it derived from recent biological production or has it already been extensively degraded? Does it represent an important N sink for the inorganic N entering the CCE system? In order to address some of these questions, I examined the quality of nitrogenous marine organic matter in water samples collected by Dr. Lihini Aluwihare's lab. Previous studies have shown that enantiomeric (D/L) ratios and non-protein amino acids serve as indicators for the "freshness" of organic matter. Our aims are to utilize these indicators to gain insight into the quality of nitrogenous organic matter in our own samples.

During my REU internship, I have been identifying the retention times of amino acids, producing standard curves for comparison with our samples, and determining the amino acid content of various water samples. My experience as an REU has given me greater insight into not only geochemical research, but also into working in an academic environment. I hope to utilize the knowledge and experience that I have gained through my internship in future graduate school and professional opportunities.
Emy Daniels

The Palenik and Brahamsha Labs at SIO study many aspects of Synechococcus, a marine bacterium which play an important role in photosynthesis throughout the world's oceans. During this past summer, I focused on identifying their grazers as part of my undergraduate research experience. One of my lab duties was to collect SIO Pier water samples each week. Some water was preserved with gluteraldehyde for later use, and some was filtered and also stored. Following the most recent annual spring bloom of synechococcus, I was able to create (from a filter sample) a library of ciliates, which are one group of known grazers of these bacteria. This method consisted of extracting all DNA from the filter and running a PCR with ciliate-specific primers to amplify sequences only from ciliate DNA. Cloning was done with E. coli to separate the different ciliate DNA which was then sent out to be sequenced. Six groups of ciliates were identified to be present following the bloom.

This molecular method of identification, without cloning, was also done on cultured nanoflagellate isolates, another group of known synechococcus grazers. I was also able to work on a method for single cell PCR which will allow individual cells to be picked out of a sample using microscopy and can compare morphological and molecular approaches to identification.
I was also able to participate in a CalCOFI-LTER research cruise during my internship. I helped to deploy nets and the CTD rosette. We took a lot of water samples and I learned how the various measurements are taken. Preserving samples and filtering water on a rocking ship was great experience. I was also able to collect some samples to compare the ciliate diversity onshore vs. offshore and surface vs. chlorophyll max. I intend to continue this project toward UCSD's BS/MS program. This REU program allowed me to experience different lab settings and sampling out at sea which helped prepare me for a future in marine biology.
Brian Pearson

For my REU internship, I performed research on phycobiliproteins, which are photosynthetic pigments that exist mainly in cyanobacteria and red algae. My specific role was in creating a method to test for the presence of phycobiliproteins in ocean samples, which was done mainly by improving upon a protocol that was used in the 1980s. This new technique was then used as a biochemical method to validate a new instrument which implements advanced laser fluorescence as an easier and faster way to test for the presence of phycobiliproteins. Because phycobiliproteins are found mostly in cyanobacteria, their presence in ocean samples denotes the presence of cyanobacteria which are an important part of marine ecology. The methods used to test for their presence are based on the unique fluorescence spectrum for each individual phyocobiliprotein. The goal of this method is to determine what phycobiliproteins are present in the ocean sample and therefore determine the present cyanobacteria communities.

As a mechanical engineering major, I did not even know what phyocobiliproteins were at the start of this REU experience; and yet with the guidance of Dr. Greg Mitchell and the excellent group of lab technicians, undergraduate 199 students, and graduate students, this has been a tremendous learning opportunity. I have learned not only a lot of biology, but how to process data, various lab techniques, and a great research experience that will continue to benefit me later in graduate school.
2007
Kate Tsyrklevich

For my REU internship, I performed research on mesozooplankton, which are a group of organisms that consume other plankton and are in the size range of approximately 0.2mm to 5mm. More specifically, I studied mesozooplankton grazing using gut pigment fluorescence as a proxy. The purpose of the study was to enhance our knowledge of nutrient cycling in the ocean by assessing the impact of zooplankton grazing on phytoplankton in various parts of the California Current as well as throughout the day-night cycle.
Samples were collected in the California Current System in May- June of 2006 and April of 2007 and frozen in liquid nitrogen in order to be accessed at a later time. Back on land at SIO, lab analysis involved microscopy for identifying organisms and picking out phytoplankton. Zooplankton were put into acetone to extract Chlorophyll pigments, which indicate how much phytoplankton was consumed, from their gut contents, and the pigment fluorescence was measured. I then learned how to organize data and analyze the results.

Overall, this experience taught me a great deal. I learned new lab techniques, data analysis strategies and everything there is to know about zooplankton grazing such as effects of temperature and food concentrations on ingestion. Weekly meetings with my mentor, Dr. Mark Ohman, provided me with a breadth of new knowledge and the opportunity to work with a distinguished faculty member. This internship provided me with a great research experience that I will be able to carry into graduate school and future job opportunities.
Randelle Bundy

As an REU, I participated on a cruise in summer 2007 to investigate the ecological role of iron in the southern California Current system (Dr. Kathy Barbeau's lab). Due to the importance of iron as an essential nutrient for phytoplankton and its extremely low abundance in seawater, the supply of iron to the California Current is vital to the ecosystem. Previous studies have focused on iron limitation in High Nutrient Low Chlorophyll (HNLC) regions of the oceans, but iron limitation is also hypothesized to play a significant role in mesotrophic (relatively low nutrient) upwelling systems like the CCE LTER study area. The cruise that I participated in was specifically investigating the influence of iron and light as co-limiting factors for phytoplankton growth in the deep chlorophyll maximum layer (DCM). The DCM is a ubiquitous feature in the offshore waters of our study area, and in the oceans worldwide.

I was able to participate in this cruise off the coast of San Diego and assist in water sampling using the CTD rosette, GO-Flos, and pole sampler. I also assisted with incubation experiments, sampling for hydrographic data, and processing samples for trace metal analysis. All of these experiences have helped me to gain a much better understanding of how oceanographic field research is conducted, and why long-term ecological research on the California Current is important for studies of climate change. I hope to continue oceanographic research in graduate school.